Scientists turn data into disease detective to predict dengue fever and malaria outbreaks
Scientists from IBM are collaborating with Johns Hopkins University and University of California, San Francisco to combat illness and infectious diseases in real-time with smarter data tools for public health. The focus is to help contain global outbreaks of dengue fever and malaria by applying the latest analytic models, computing technology and mathematical skills on an open-source framework.
Vector-borne diseases, like malaria and dengue fever, are infections transmitted to humans and other animals by blood-feeding insects, such as mosquitoes, ticks and fleas.
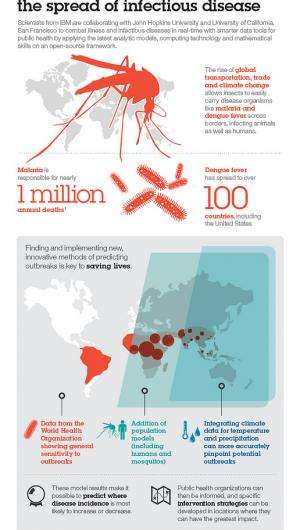
Once thought to be limited geographically to the tropics or developing countries, they continue to show up all over the world and are among the most complex and dangerous infectious diseases to prevent and control. The rise of global transportation, trade and climate change allows insects to easily carry disease organisms across borders, infecting animals as well as humans. Dengue fever, for example, has spread to over 100 countries, including the United States, and malaria is responsible for over one million annual deaths. Finding and implementing new, innovative methods of predicting outbreaks is key to saving lives.
Epidemiologists rely on disease and vaccine simulations to determine the spread of global infection. Until recently, these models were hosted on closed systems and took years to produce due to inefficient data collection and lack of computing power. This approach to model development makes it too slow to respond on a timescale relevant to unexpected pandemics as large populations can be crippled by never before seen viruses or illnesses in a matter of days or weeks. Scientists need to understand not only the dynamics of the disease itself, but also the spread of insect vectors and contributing environmental factors.
Using existing vector borne disease models from Johns Hopkins and UCSF, IBM researchers are developing new dengue fever and malaria models that are shared as part of the open-source modeling application, Spatio Temporal Epidemiological Modeler (STEM). The scientists identified the opportunity to combine population analytics, algorithms of disease paths and powerful computing to build realistic and accessible models of these infectious diseases. This capability allows for the study of disease dynamics in humans and intervention strategies such as vector control and vaccine distribution.
"Public health officials can't afford to act on speculation during an epidemic. They need accurate and timely access to data to see what the potential spread of a disease might be for a given geographic region over a period of time,' said James Kaufman, Public Health Manager, IBM Research. 'The scientific and research communities are collaborating to lead a new age of science-based, data-centric disease modeling to protect the health of people. By understanding how and why these diseases spread, we can identify those regions most susceptible to emerging disease, inform public health, and allow them to focus on specific interventions in locations where they can have the greatest impact."
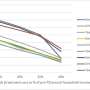
In the case of malaria, using both the model and data from the World Health Organization, IBM and Johns Hopkins demonstrated new analytic measures for the sensitivity of malaria incidence to changes in local climate factors such as temperature and precipitation. Knowing this sensitivity makes it possible to predict where malaria incidence is most likely to increase or decrease based on predicted changes in local weather and environmental conditions in a specific region.
"There are a lot of tacit assumptions out there about how changes in climate will impact the distribution of diseases like malaria. This work suggests that things probably are not so simple, a change that has a huge effect on malaria transmission in one place might not be as important somewhere else," said Justin Lessler, Johns Hopkins Bloomberg School of Public Health. "One of the nice things about open source projects like STEM is that now whoever wants to can download the model and start tweaking it, seeing if their own data or assumptions fundamentally change the results."
Previous models of dengue fever treated the mosquito vector indirectly, approximating transmission as a human-to-human contact process. IBM Research and UCSF used STEM's ability to build models on top of models and integrate them with location-specific climate data. This allowed to the inclusion of the vector population into existing models, providing a more realistic description of the disease dynamics, which can present public health officials more effective predictions of epidemics spread.
"It is important to recognize the synergistic effort of theoretical and computational scientists, disease experts and public health officials making a difference in how rapidly and effectively we fight infectious diseases," said Simone Bianco, UC San Francisco, Bioengineering and Therapeutic Sciences. "We have to be ready at the drop of a hat to parse through disparate data from global disease surveillance systems, conduct computationally intense research and transfer our knowledge to public health officials to help them visualize population health, detect outbreaks, develop new models, and evaluate the effectiveness of policies".
Available through the Eclipse Foundation, STEM is free and open to any scientist or researcher who chooses to build on and contribute to its library of models, computer code and denominator data. This openness facilitates the development of advanced mathematical models, the creation of flexible models involving multiple populations (species), interactions between diseases, and a better understanding of epidemiology.
These achievements were published in the peer-reviewed journals Malaria and Theoretical Biology.