Technique for analyzing specific T-cell responses could pave way for more rationally designed vaccines
Among different populations, a vaccine can vary in efficacy against a specific pathogen. Unraveling the discrepancy requires tools to analyze the exact nature of responses by a specific type of immune cell: the T cell. Now, a research team from Singapore and the United States has developed an approach that simultaneously identifies and characterizes T cells specific to a variety of antigens—the part of the pathogen recognized by the immune system. The work could lead to more personalized and thus effective vaccine designs.
"[Our] approach can be applied to the study of any vaccine that elicits a T-cell response, allowing for an in-depth analysis of the breadth and quality of the antigen-specific T-cell response," says Evan Newell of the A*STAR Singapore Immunology Network, who led the study.
The new method involves a combination of two technologies. First, complexes of candidate peptide antigens and major histocompatibility complex (MHC) molecules, which bind antigens for presentation to T cells, are each tagged with a unique combination of heavy metal markers. These so-called 'tetramers' are then incubated with patient samples containing T cells, which attach to any complexes with peptides that match those recognized by receptors on the T-cell surface. Next, a high-throughput, multiplexing technique called mass cytometry is used to analyze the coupled T cells at single-cell resolution. "This yields an unprecedented amount of information from each sample," says Newell.
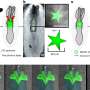
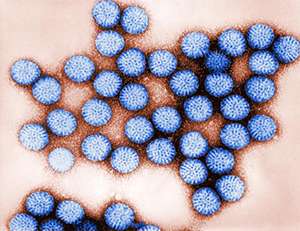
As a proof of principle, Newell and his co-workers analyzed blood and intestinal tissue from healthy volunteers for T cells that recognize rotavirus (see image). "Rotavirus infection is a major problem in the developing world, partially because vaccines do not work well there," explains Newell. The gut pathogen causes severe diarrhea in young children and kills more than 500,000 people worldwide each year.
The researchers stained the T cells from the healthy individuals with a series of 77 candidate rotavirus epitopes. Each epitope is a slightly different site on a rotavirus protein—the antigen—that is recognized by the immune system. From this larger collection, they identified and validated six rotavirus epitopes and simultaneously analyzed the T cells specific for each of them.
"We were surprised to find that T cells targeting different parts of the rotavirus had dramatically different characteristics; and, only one set of these T cells were found to be present in the intestine of normal donors," says Newell. "These analyses should allow faster identification of correlates for vaccine efficacy that can be used to speed the testing of new vaccines," he adds.
More information: Newell, E. et al. Combinatorial tetramer staining and mass cytometry analysis facilitate T-cell epitope mapping and characterization, Nature Biotechnology 31, 623–629 (2013). www.nature.com/nbt/journal/v31 … 7/full/nbt.2593.html