IRB develops ChroGPS, a new generation visual browser of the epigenome
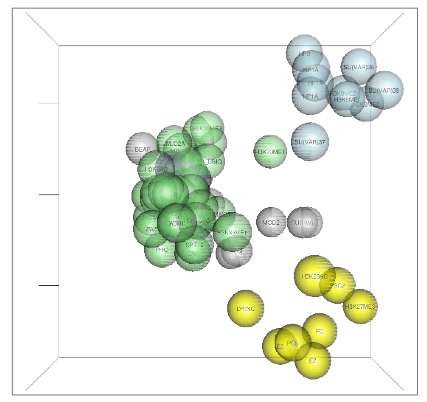
ChroGPS is a software application that serves to facilitate the analysis and understanding of epigenetic data and to extract intelligible information, which can be downloaded free of charge in Bioconductor, a reference repository for biocomputational software. The scientists at the Institute for Research in Biomedicine (IRB Barcelona) describe the uses of the programme in an article published in the journal Nucleic Acids Research, in which they explain that ChroGPS is the answer to a problem that has been dragging on for the last ten years.
In the last 15 years, researchers worldwide have generated a large amount of information about the epigenome: proteins, factors and epigenetic markers which, when bound to DNA, regulate gene expression. Enormous projects such as ENCODE (for humans and mice) or modENCODE (for other lab model systems, such as the fly Drosophila or the worm C. elegans) have been devoted to collecting these data in order to analyse and interpret them in the framework of genomic data and to form hypotheses about functions and relations. In spite of these efforts, tools are still needed to extract functional and relational information about the epigenome and to present the results in a visual manner, as ChroGPS does.
"With ChroGPS we wanted to integrate epigenetic data with genetic data to reap the great benefits from them and to be able to understand this information. The analyses continue to be extremely complex and the results to be interpreted very unclear," says Ferran Azorín, head of the Chromatin Structure and Function lab at IRB Barcelona and CSIC researcher professor, who studies epigenomic regulation. "With this tool we have reached the same conclusions as those presented in Nature by researchers working on the modENCODE, but the enormous difference is that instead of seeing the information in hundreds of graphs and figures like in modENCODE, we have achieved a single map," explains Azorín.
The initiative emerged from dialogue between Azorín's group, through the PhD student Joan Font-Burgada, and the bioinformatician Òscar Reina, a member of the Biostatistics and Bioinformatics Unit of IRB Barcelona, which was managed by David Rossell at that time. "ChroGPS is based on the sequential application of two steps: first the generation of distances (or degrees of similarity) between epigenetic components on the basis of several possible measurements that we have developed, and after, in the representation of these distances in the form of bi- o tri-dimensional maps to facilitate their interpretation. For example, they are like visual maps from which distance tables can be drawn up in kilometers between cities," describes Òscar Reina, one of the developers of the software application. "The most important thing for us in this first stage has been to present the biological information in a simple but at the same time reliable manner from the point of view of data treatment, for example correcting systematic biases between experiments that can lead to erroneous conclusions," adds Rossell, who is now at the University of Warwick, in the UK.
Now that the programme is available to the entire community, the researchers contemplate new challenges with ChroGPS. Among his objectives, Ferran Azorín aims to follow the complex transformation of a healthy cell into a cancerous one through tracking the genetic and epigenetic changes that occur. To tackle this project with ChroGPS, the researchers will have to take new steps in statistical and mathematic methods.
More information: "chroGPS, a global chromatin positioning system for the functional analysis and visualization of the epigenome." Joan Font-Burgada, Oscar Reina, David Rossell, and Fernando Azorín. Nucleic Acids Research (2013) DOI: 10.1093/nar/gkt1186