Team finds mutations expressed within melanoma tumors that predict effective responses to a groundbreaking immunotherapy
A team led by Ludwig and Memorial Sloan Kettering (MSK) researchers has published a landmark study on the genetic basis of response to a powerful cancer therapy known as immune checkpoint blockade. Their paper, in the current issue of the New England Journal of Medicine, describes the precise genetic signatures in melanoma tumors that determine whether a patient will respond to one such therapy. It also explains in exquisite detail how those genetic profiles translate into subtle molecular changes that enable the immune system attack of cancer cells in response to immune checkpoint blockade.
"The genetic signature we have found will be invaluable to understanding the biological mechanisms that drive therapeutic responses to immunotherapy for metastatic melanoma," says Jedd Wolchok, MD, PhD, director of the Ludwig Collaborative Laboratory and associate director of the Ludwig Center for Cancer Immunotherapy at MSK, who co-led the study with Timothy Chan, MD, PhD, of MSK's Human Oncology and Pathogenesis Program. "Further, our strategy can now be applied to determine the genetic signatures associated with the efficacy of a number of other immunotherapies and cancers."
Few approaches to treating cancer have generated as much excitement as immunotherapy, in which the immune system is engaged to destroy malignancies. One class of such treatments targets CTLA-4, a molecule expressed on the surface of killer T cells that ordinarily blocks their proliferation. Antibody drugs that block CTLA-4 thus stimulate killer T cell responses—which can target cancer cells—and significantly extend survival for many melanoma patients. Yet not all patients respond equally to this treatment: some, remarkably, survive many years; others fail to respond at all.
"There is a subset of melanoma patients who are living far longer than anyone would have expected in the past, largely because of this treatment and other recently developed targeted and immunologic treatments," says Wolchok. "But we did not know how to identify them, and that's what really drove this investigation."
Cancer cells are swift but sloppy proliferators, generating countless mutations across their genome as they multiply. Those mutations are often expressed as changes in the chains of amino acids that make protein molecules. Like all cells, cancer cells chop up and hold out short fragments of such proteins—each about 9 amino acids in length—for the immune system to assess. These "peptides" are held up and presented to immune cells by a protein complex known as MHC Class I, which varies significantly between people.
"Previous studies by Jedd and others had shown that the particular MHC type of a patient doesn't appear to influence the efficacy of CTLA-4 blockade," says Chan. "So we decided to see if the tumor genome has anything to say about whether or not people respond to this therapy. The result was entirely unexpected, and the answer is exceedingly important."
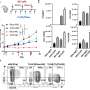
Chan, Wolchok and their colleagues initially hypothesized that tumors that harbored highly mutated cells would be most responsive to CTLA-4 blockade. To test that hypothesis, they sequenced and compared all of the genes expressed as proteins (collectively known as the "exome") in tumors taken from 25 patients treated with anti-CTLA-4 antibodies and found that this was, to some degree, true. "But looking at the data a little more deeply," says Wolchok, "we saw that there were outliers—patients who had over one thousand mutations who didn't respond, and some with just a few dozen who did. This was a strong indication that the quality of the mutations matters."
A sophisticated computational analysis of the cancer genomes revealed that a set of core peptide sequences—each four amino acids long (tetrapeptides)—within MHC Class I-presented peptides were unequivocally associated with response to treatment. To test the prognostic power of this genetic signature, the researchers sequenced the exomes of tumors from another 39 melanoma patients treated with CTLA-4 blockade. They found that all those in this set who had responded to the therapy had at least one and typically several more of the tetrapeptides they had identified. Those who failed to respond did not. Their results show that the mutant DNA sequences, can occur anywhere in the genome—not just within mutant "driver" genes that are already known to contribute to cancer.
"The more mutated the tumor's genome is," says Chan, "the more likely it is that immunotherapy will work. Since tumors induced by tobacco—such as those of non-small cell lung cancer—have more mutations than most other cancers except melanoma, this finding has enormous medical implications for these genetically diverse cancers."
It also helps explain, says Wolchok, why the relatively more mutated cancers have been found in clinical trials to be the most responsive to checkpoint blockade.
The researchers further validated their findings by showing in lab experiments that killer T cells taken from responsive patients were potently activated by synthetic replicas of the mutated tetrapeptides, but not by their normal counterparts. T cells from healthy people failed to respond to mutant peptides, indicating that the T cell response in question was specific to melanoma.
Examination of the mutations indicates that the ones that induce clinically relevant immune responses do so in two major ways. Some bolster the presentation of the peptides by HLA molecules; others generate changes that killer T cells "see" anew as signs of a lurking danger. Some of the mutations, it turns out, change the peptides in a way that make them identical to protein fragments associated with infectious bacteria and viruses.
The researchers will next perform similar analyses on other immunotherapies and types of cancer, most notably lung cancer.