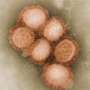
Tracing evolution of chicken flu virus yields insight into origins of deadly H7N9 strain
An international research team has shown how changes in a flu virus that has plagued Chinese poultry farms for decades helped create the novel avian H7N9 influenza A virus that has sickened more than 375 people since 2013. The research appears in the current online early edition of the scientific journal Proceedings of the National Academy of Sciences.
The results underscore the need for continued surveillance of flu viruses circulating on poultry farms and identified changes in the H9N2 virus that could serve as an early warning sign of emerging flu viruses with the potential to trigger a pandemic and global health emergency. The work focused on the H9N2 chicken virus, which causes egg production to drop and leaves chickens vulnerable to deadly co-infections. Scientists at St. Jude Children's Research Hospital and the China Agricultural University, Beijing, led the study.
Researchers used whole genome sequencing to track the evolution of the H9N2 chicken virus between 1994 and 2013. The analysis involved thousands of viral sequences and showed that the genetic diversity of H9N2 viruses fell sharply in 2009. From 2010 through 2013 an H9N2 virus emerged as the predominant subtype thanks to its genetic makeup that allowed it to flourish despite widespread vaccination of chickens against H9N2 viruses.
Evidence in this study suggests the eruptions set the stage for the emergence of the H7N9 avian virus that has caused two outbreaks in humans since 2013, with 115 confirmed deaths. The H9N2 infected chickens likely served as the mixing vessel where H9N2 and other avian flu viruses from migratory birds and domestic ducks swapped genes, researchers noted. The resulting H7N9 virus included six genes from the H9N2.
"Sequencing the viral genome allowed us to track how H9N2 evolved across time and geography to contribute to the H7N9 virus that emerged as a threat to human health in 2013," said Robert Webster, Ph.D., a member of the St. Jude Department of Infectious Diseases. He and Jinhua Liu, Ph.D., of the College of Veterinary Medicine at the China Agricultural University, are co-corresponding authors.
"The insights gained from this collaboration suggest that tracking genetic diversity of H9N2 on poultry farms could provide an early warning of emerging viruses with the potential to spark a pandemic," Webster said.
The analysis also provided insight into the creation of the H9N2 virus that emerged as the predominant subtype in 2010. Factors included widespread use of poultry vaccines and the natural tendency of flu to mutate, mix and swap genes.
Beginning in 1998, vaccinating poultry against H9N2 prevented flu outbreak for more than a decade. Vaccines work by recognizing and attaching to the spike-shaped hemagglutinin (HA) protein on the surface of the flu virus. That blocks the virus from infecting healthy cells. Changes in the HA gene that change the shape of the HA protein can reduce vaccine effectiveness and result in disease outbreaks. HA mutations occur naturally over time. Vaccines increase pressure for HA mutations that help the virus escape vaccine detection and cause infection.
Researchers at the China Agricultural University checked H9N2 vaccine effectiveness against the predominant H9N2 virus from 2010-11. Working in vaccinated and unvaccinated chickens, investigators found the vaccine neither protected vaccinated chickens from infection nor prevented spread of the virus in vaccinated chickens. Those failures suggest that due to HA mutations vaccines were less able to recognize the virus.
The tendency of flu viruses to swap genes also contributed to the enhanced ability of the predominant H9N2 subtype to spread. Researchers found that prior to the virus' emergence as the predominant H9N2 the virus had swapped genes with quail and duck influenza viruses.
The combination fueled the recent outbreaks of H9N2 on chicken farms by helping the virus escape vaccine detection and spread rapidly in vaccinated and unvaccinated poultry, said co-first author Juan Pu, Ph.D., a St. Jude visiting scientist from the China Agricultural University. The other first authors are Shuoguo Wang, Ph.D., of the St. Jude Department of Computational Biology, and Yanbo Yin, Ph.D., of Qingdao Agricultural University, Qingdao, China.
"The emergence of this dominant H9N2 virus was the first step in the genesis of the H7N9 viruses because it greatly increased the likelihood of reassortment between H9N2 and other flu subtypes," Liu said. Reassortment refers to the tendency of flu viruses to swap genes.
More information: Evolution of the H9N2 influenza genotype that facilitated the genesis of the novel H7N9 virus, PNAS, www.pnas.org/cgi/doi/10.1073/pnas.1422456112