Study explains how dengue virus adapts as it travels, increasing chances for outbreaks
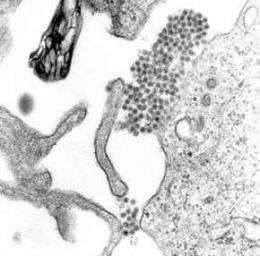
A researcher from The University of Texas Medical Branch at Galveston is an integral member of a collaborative group that is the first to explain the mechanisms that the Dengue virus has developed to optimize its ability to cause outbreaks as it travels across the globe to new places and revisits old ones. An early online version of this paper detailing the findings has recently been published in Science.
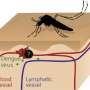
Dengue virus has been spreading throughout warm regions of the world, prompting the virus to adapt to new environments. This diversification in viral strains has resulted in the development of strains that appear associated with greater potential for sparking epidemics. Several dengue outbreaks have occurred when new dengue strains emerged and displaced the native strains that the local population had already developed immunity against. Until now, the mechanisms governing how and why some viral strains are better suited for causing widespread disease has been poorly understood.
The investigators examined the different clades of dengue virus-2 known to be circulating around Puerto Rico in 1994 when a severe epidemic broke out. Investigating the differences between the virus strain that was most commonly seen from 1986 to 1995 and a new, more potent viral strain that was first isolated in 1994 was the key to figuring out why this outbreak occurred.
They identified an interaction between the newcomer virus's RNA and proteins within the host that allows the virus to bypass the host's immune response, making it easier for the virus to invade. Based on the findings, the research team devised a model to explain the 1994 dengue outbreak in Puerto Rico.
"This study highlights the critical and oft forgotten role played by non-coding RNAs in the battle between viruses and their human hosts," said author Mariano Garcia-Blanco, UTMB professor and chair of the department of biochemistry and molecular biology and also professor of emerging infectious diseases at the Duke-NUS Graduate Medical School in Singapore. "It emphasizes the importance of multidisciplinary research: a fabulous marriage of basic RNA biology and clinically informed epidemiology uncovered an unexpected route of virus evolution that explained (and perhaps could predict) epidemic potential."
More information: Dengue subgenomic RNA binds TRIM25 to inhibit interferon expression for epidemiological fitness, Science DOI: 10.1126/science.aab3369