Scientists find 1500 'ageing' genes that could lead to new treatments
Researchers from The University of Queensland have helped identify nearly 1,500 genes associated with ageing that could lead to new health treatments.
Dr Joseph Powell, from UQ's Institute for Molecular Bioscience (IMB), said the discovery, made by an international team of scientists, could lead to improved prevention and treatment for age-related diseases.
"We examined the link between gene activity and ageing, identifying genes that became more or less active as people got older," Dr Powell said.
"This information could be used to predict people who are at risk relative to their age for disease, and allow people to make lifestyle and environmental changes to mitigate that risk.
"While an anti-ageing treatment is still well into the future, this work could provide a roadmap for developing new treatments for age-related diseases."
The team made the discovery by studying gene activity levels in the blood of over 15,000 people, pinpointing 1,497 age-associated genes, most of which were previously either not known or not linked with ageing.
The UQ researchers, including Dr Powell and Professor Peter Visscher from UQ's Queensland Brain Institute, used this information to develop a new method of predicting biological age based on gene activity profiles.
"Participants with a higher biological age, meaning their predicted age based on their gene activity profile was higher than their actual chronological age, also had worse disease risk profiles, for example, increased blood pressure and cholesterol levels," Dr Powell said.
"Conversely, some participants had a lower biological age, indicating they may be healthier than others of the same chronological age."
The method has been incorporated into a website where researchers analysing human gene activity data can predict the biological age of their participants, and assess whether participants are ageing faster or slower than expected.
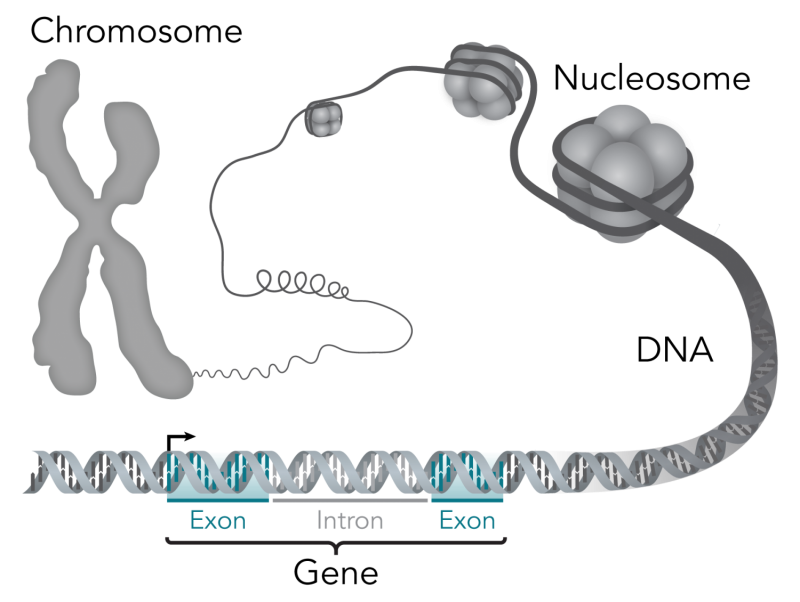
The study also identified that the changes in gene activity levels that occur as we age appear in conjunction with changes to the enhancer regions of DNA, which can act as dimmer switches to regulate the activity of genes.
"This study has illuminated some of the underlying molecular mechanisms that cause your disease susceptibility to rise as you age, opening new avenues for future studies," Dr Powell said.
More information: Marjolein J. Peters et al. The transcriptional landscape of age in human peripheral blood, Nature Communications (2015). DOI: 10.1038/ncomms9570