Research team finds that protein motions regulate HIV infectivity
Nearly 37 million people worldwide are living with HIV. When the virus destroys so many immune cells that the body can't fight off infection, AIDS will develop. The disease took the lives of more than a million people last year.
For the past three and a half years, a team of researchers from six universities, led by the University of Delaware and funded by the National Institutes of Health and the National Science Foundation, has been working to uncover new information about a protein that regulates HIV's capability to hijack a cell and start replicating. Their findings, reported recently in the Proceedings of the National Academy of Sciences, point to a new avenue for developing potential strategies to thwart the virus.
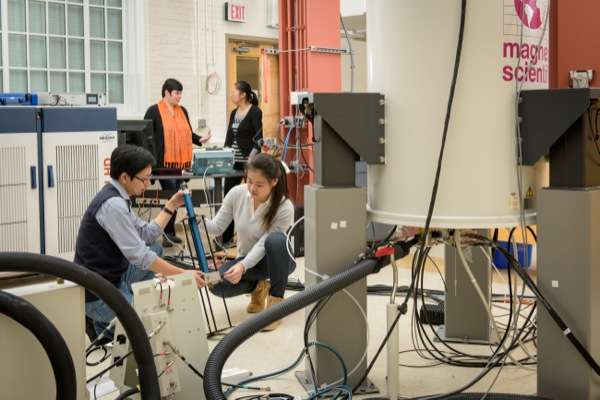
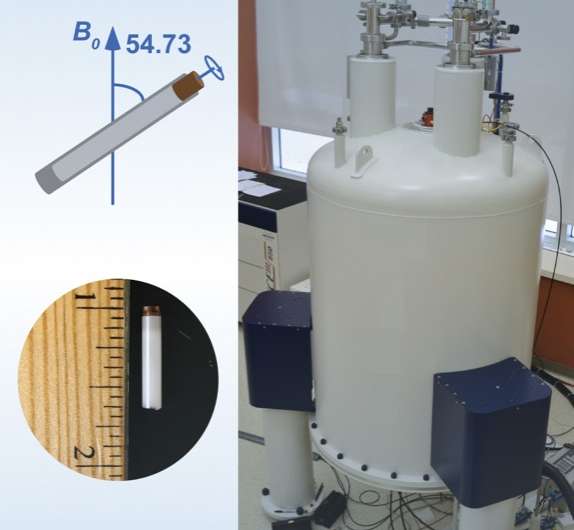
The team included scientists from UD, the University of Pittsburgh School of Medicine, University of Illinois at Urbana-Champaign, Carnegie Mellon University, the National High Magnetic Field Laboratory at Florida State University and Vanderbilt University School of Medicine. They used a combination of high-tech tools and techniques, including magic-angle-spinning nuclear magnetic resonance (NMR) spectroscopy and computer simulations of molecules, to examine the interactions between HIV and the host-cell protein cyclophilin A (CypA), right down to the movement of individual atoms.
"In a nutshell, we found that the infectivity of HIV is regulated by the motions of these proteins," says Tatyana Polenova, professor of chemistry and biochemistry at the University of Delaware, who led the study. "It's a subtle regulation strategy that does not involve major structural changes in the virus."
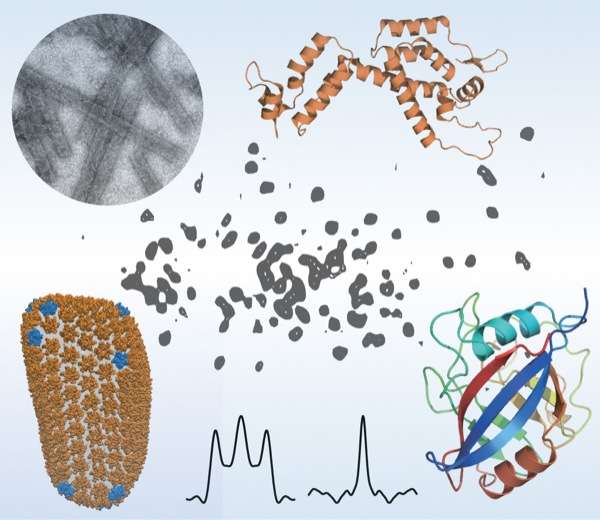
Sixty times smaller than a red blood cell, HIV contains a cone-shaped shell, or capsid, made of protein, which surrounds two strands of RNA and the enzymes the virus needs for replication. Like any virus, HIV can only produce copies of itself once it has invaded a host organism. Then it will begin directing certain host cells to begin producing the virus.
But how does HIV invade a cell? In humans, the protein CypA can either promote or inhibit viral infection through interactions with the HIV capsid, although the exact mechanism is not yet known. A portion of the HIV capsid protein, called the CypA loop, is responsible for binding to the CypA in the human host cell. Once this occurs, the virus typically becomes infectious.

However, a change of just one amino acid in the CypA loop can cause the virus to operate opposite from how it does normally, allowing the virus to become non-infectious when CypA is present, and to become infectious when there is no CypA present. Such changes are called "escape mutations," Polenova says, because they allow the virus to "escape" from its dependence on CypA.
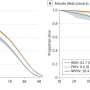
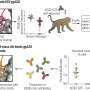
To home in on this escape mechanism, the research team examined assemblies of different variants of HIV capsid protein complexed with CypA. Using magic-angle-spinning NMR, they recorded the motions in these assemblies, atom by atom, on time scales ranging from nanoseconds to milliseconds, from a billionth of a second to a thousandth of a second.
The team found that a reduction in the naturally occurring motions in the binding region due to the mutations allowed the virus to escape from CypA dependence. Magic-angle-spinning NMR experiments provided a direct probe of these motions, recording the changes in the magnetic interactions between nuclei. Computer simulations allowed the team to visualize the motions.
Some portions of the capsid protein do not move at all or move only a little while other portions undergo large-amplitude motions distributed over a wide range of time scales, with the most dynamic region being the CypA loop. Polenova says it is rather surprising that such extensive motions are present in the assembled capsid, and that these dynamics could be detected by both NMR and computer simulations.
"It is the first time that quantitative agreement between experiment and computation was achieved in a dynamics study, and it's particularly exciting that this was attained for such a complex system," Polenova says. "We hope this work may guide the development of new therapeutic interventions, such as small molecules that would serve as interactors with the HIV capsid and inhibit these dynamics."

Polenova says the diverse team of researchers, with expertise in HIV virology, structural biology, biophysics and biochemistry, was critical to the study's success, along with access to national high-field NMR facilities through the National High Magnetic Field Laboratory. The team was assembled through the NIH-funded Pittsburgh Center for HIV Protein Interactions. Led by Prof. Angela Gronenborn, the center brings together high-caliber scientists and facilities to elucidate the interactions of HIV proteins with host cell factors.
The research team at UD included doctoral students Manman Lu, Changmiao Guo and Christopher L. Suiter, NMR spectroscopist Guangjin Hou, and postdoctoral researcher Huilan Zhang. Prof. Angela M. Gronenborn led the team from the Pittsburgh Center for HIV Protein Interactions and the Department of Structural Biology at the University of Pittsburgh School of Medicine, which included Jinwoo Ahn, Jn-Ja L. Byeon and Peijun Zhang. Also collaborating on the project were Christopher J. Langmead from Carnegie Mellon University, Ivan Hung, Peter L. Gor´kov, Zhehong Gan and William Brey from the National High Magnetic Field Laboratory at Florida State University, Christopher Aiken from Vanderbilt University School of Medicine, and Juan Perilla and Klaus Schulten from the University of Illinois.
The research and the scientific instrumentation used were supported by the National Institutes of Health and the National Science Foundation. Work performed at the National High Magnetic Field Laboratory also was co-supported by the State of Florida.
Polenova and her research team published a second article, reporting on the first atomic resolution structure of a protein bound to polymerized microtubules, in the same issue of the Proceedings of the National Academy of Sciences.
More information: Manman Lu et al. Dynamic allostery governs cyclophilin A–HIV capsid interplay, Proceedings of the National Academy of Sciences (2015). DOI: 10.1073/pnas.1516920112