A cultural revolution in the study of the gut microbiome
It's estimated that as many as 1,000,000 Americans suffer from inflammatory bowel diseases (IBD), such as ulcerative colitis and Crohn's disease, which cause mild to severe symptoms that at best can be managed and at worst can lead to life-threatening complications. While abnormal immune responses are largely responsible for these diseases, issues relating to gut microbiome, intestinal epithelial cells, immune components and the gut's rhythmic peristalsis motions can also contribute to and exacerbate symptoms. But until now, scientists have been hard-pressed to develop new therapies for treating IBDs due to their inability to replicate the human gut microenvironment in the laboratory.
Now, a team at the Wyss Institute for Biologically Inspired Engineering at Harvard University co-led by Wyss Institute Founding Director Donald Ingber, M.D., Ph.D., and Wyss Core Faculty member James Collins, Ph.D., has leveraged the Institute's proprietary human-organs-on-chips technology to microengineer a model of human intestinal inflammation and bacterial overgrowth in a human-gut-on-a-chip. The advance, reported in Proceedings of the National Academy of Sciences (PNAS) journal, for the first time allows scientists to analyze how normal gut microbes and pathogenic bacteria contribute to immune responses, and to investigate IBD mechanisms in a controlled model that recapitulates human intestinal physiology.
"Chronic inflammation of the intestine is thought to be caused by abnormal interactions between gut microbes, intestinal epithelial cells and the immune system, but so far it has been impossible to determine how each of these factors contribute to the development of intestinal bowel disease," said Hyun Jung Kim, Ph.D., former Wyss Technology Development Fellow and first author on the study, speaking about the limitations of conventional in vitro and animal models of bacterial overgrowth and inflammation of the intestines.
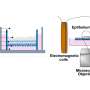
The human gut-on-a-chip technology, however, provides an ideal microenvironment for mimicking the natural conditions of the human intestines in a small-scaled, controllable in vitro platform. The human gut-on-a-chip was first invented at the Wyss Institute in 2012. Made of a clear flexible polymer about the size of a computer memory stick, the hollow-channeled microfluidic device simulates the physical structure, microenvironment, peristalsis-like motion waves and fluid flow of the human intestine.
In this latest advance reported in PNAS, the Wyss team showed that the human gut-on-a-chip's unique ability to co-culture intestinal cells with living microbes from the normal gut microbiome for an extended period of time, up to two weeks, could allow breakthrough insights into how the microbial communities that flourish inside our GI tracts contribute to human health and disease.
"The discovery of the microbiome and its significance represents a huge paradigm shift in our understanding of human health - there are more microbes living on us and in us than our own cells," said Ingber, who is also the Judah Folkman Professor of Vascular Biology at Harvard Medical School and the Vascular Biology Program at Boston Children's Hospital, and Professor of Bioengineering at the Harvard John A. Paulson School of Engineering and Applied Science. "Until now, use of traditional culture methods and even more sophisticated organoid cultures have prevented the microbiome from being studied beyond one or two days. With our human gut-on-a-chip, we can not only culture the normal gut microbiome for extended times, but we can also analyze contributions of pathogens, immune cells, and vascular and lymphatic endothelium, as well as model specific diseases to understand complex pathophysiological responses of the intestinal tract."
"There is much to be learned about IBD, as well as how antibiotics impact the microbiome," said Collins, who is also Termeer Profesor of Medical Engineering and Science at the Massachusetts Institute of Technology. "This technology enables one to study in an isolated and controlled manner the complexity of the microbiome and the role different microbial species play in health and disease. It is therefore a highly valuable platform for discovery and clinical translation efforts."
Already the advance has revealed new discoveries into the inner workings of the human intestinal tract and its immune responses. Four small proteins that stimulate inflammation (called cytokines) were found to work in tandem to trigger inflammatory immune responses that damage and irritate the bowel. This discovery could open a new potential therapeutic pathway to treating IBD by "blocking" these cytokine proteins simultaneously.
The Wyss team also studied the role fluid flow and the wave-like peristaltic movement of the gut plays in maintaining a dynamic equilibrium of the gut microbiome, finding that absence of peristaltic movement can lead to rampant overgrowth of bacteria completely independent of changes in fluid flow. This could help explain why some patients with IBD and other conditions develop bacterial overgrowth, such as patients who develop ileus, which is a syndrome that can occur following intestinal surgery when there is a prolonged delay in the body's ability to resume normal peristaltic motions.
The Wyss team believes the ability of the human gut-on-a-chip to culture the microbiome with human gut cells also holds promise for the field of precision medicine, where a patient's own cells and gut microbiota could one day be cultured inside a gut-on-a-chip for testing different therapies and identifying an individualized treatment strategy.
"Previously the microbiome and its role in human health were largely defined through study of their gene expression, but now, by being able to carry out human experimentation in vitro relating to how the microbiome, human intestinal cells and human immune components interplay, we hope to gain a much deeper understanding of underlying pathophysiological mechanisms that will hopefully lead to development of new and more effective therapies," said Ingber.
More information: Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip, Proceedings of the National Academy of Sciences, www.pnas.org/cgi/doi/10.1073/pnas.1522193112