New research could personalize medicine for arthritis patients

Joint injury can lead to post-traumatic osteoarthritis (PTOA). In fact, about half of all people who rupture the anterior cruciate ligament (ACL) in their knee will develop PTOA within 10-20 years of the injury.
But the molecular and cellular mechanisms leading to cartilage degeneration or PTOA due to trauma are not well understood.
Recently, a team of scientists from Lawrence Livermore National Laboratory (LLNL), University of California, Davis, University of California, Merced and Regeneron Pharmaceuticals examined the whole-joint gene expression by RNA sequencing at one day, one, six and 12 weeks after injury. The team used a new, non-invasive tibial compression mouse model of PTOA, that mimics ACL rupture in humans from a single high-impact injury.
The research appears in the online edition of the Journal of Orthopaedic Research.
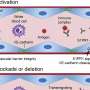
Sometimes called degenerative joint disease or "wear and tear" arthritis, osteoarthritis (OA) is the most common chronic condition of the joints. It occurs when the cartilage or cushion between joints breaks down leading to pain, stiffness and swelling.
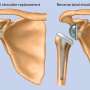
Many individuals developing OA show no signs until significant joint damage has occurred. At that point, the only available long term treatment options are surgical replacement of the joint and/or pain management.
Identifying and characterizing OA biomarkers for detecting and tracking the progression of the disease, combined with developing new pharmacologic interventions aimed to minimize cartilage damage, could personalize medical treatment before the disease is all consuming. Most importantly, treatments could be developed that, when administered immediately post injury, would prevent the development of PTOA years later.
"The goal of the study was to see if there are biomarkers associated with cartilage degradation, which could then be further explored as therapeutic targets in future experiments," said Jiun Chang, a UC Merced graduate student mentored by LLNL's Gaby Loots and the lead author of the study.
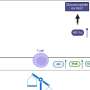
The study identified 1,446 genes differentially expressed in injured joints, including several known regulators of OA, as well as many new genes. The team also identified 18 long, noncoding RNAs differentially expressed in the injured joints, RNAs that have not yet been explored functionally in this context.
"This study provides the first account of gene expression changes associated with PTOA development and progression in this tibial compression model," said Aimy Sebastian, also a UC Merced graduate student mentored by Loots, who co-lead the study with Chang.
The research team also included LLNL staff member Deepa Murugesh; UC Davis professor Blaine Christiansen; and Sarah Hatsell and Aris Economides of Regeneron.
"By comparing our data to gene-expression data generated using the surgical destabilization of the medial meniscus PTOA model, we identified several common genes and shared mechanisms. Our study highlights several differences between these two models and suggests that the tibial compression model may be a more rapidly progressing model of PTOA," said Loots, an LLNL biologist who leads the team.
This study provides the first account of whole genome expression profiles to obtain new insights into the temporal progression of the disease.
More information: Journal of Orthopaedic Research, onlinelibrary.wiley.com/doi/10.1002/jor.23263/epdf