Liver tissue model accurately replicates hepatocyte metabolism, response to toxins
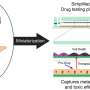
A team of researchers from the Massachusetts General Hospital (MGH) Center for Engineering in Medicine (MGH-CEM) have created a "liver on a chip," a model of liver tissue that replicates the metabolic variations found throughout the organ and more accurately reflects the distinctive patterns of liver damage caused by exposure to environmental toxins, including pharmaceutical overdose. Their report has been published online in the journal Scientific Reports.
"Our goal with this project was to create a liver tissue construct that responds to toxins the same way the liver in your body does," says William McCarty, PhD, a postdoctoral fellow at MGH-CEM and the paper's lead author. "The liver is a chemical processing plant, but it's not a single vat; different locations within the liver react differently to drugs and toxins. Here, we exploited microfluidics to control the metabolism of liver cells down to a resolution of a few cells, allowing us to create liver tissue that shows the same patterns of toxicity caused by differences in drug metabolism as the liver in your body."
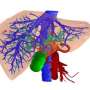
When blood passes through the liver, it travels from arteries to veins through channels called sinusoids, lined with the liver cells called hepatocytes. From one end of the sinusoid to another, the hepatocytes have different metabolic functions, often controlled by external factors and gene expression. For example, the cells closest to the arterial end of the sinusoid are most efficient at releasing glucose that has been stored in the form of glycogen, while cells at the venous end are most efficient at taking up and storing glucose. Similar differences for other liver functions are well known, with metabolic changes occurring across the 25-cell length of the sinusoid.
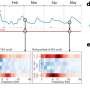
In order to develop a system that more closely replicates the metabolic differences among hepatocytes, the research team developed a microfluidic device that distributes hormones or other chemical agents across a 20- to 40-cell-wide sample of hepatocytes in such a way that the effects on the liver cells vary from one side to the other. For example, if blood-sugar-lowering insulin is fed into one of the device's two inlets while glucagon, which raises blood sugar, is added through the other, the metabolism of the hepatocytes is changed so that those on one side release glucose while those on the other take it up. The use of other agents produced similar results across the field of hepatocytes regarding nitrogen metabolism or alcohol degradation, and use of a molecule that induces the expression of drug metabolism enzymes resulted in varied zones of susceptibility to the toxic effects of acetaminophen.
"Investigators have been developing in-vitro liver models for 40 years, but all of those systems ignore the distinct patterns of metabolically active hepatocytes that exist within the liver sinusoid" says Martin Yarmush, MD, PhD, director of the MGH-CEM and the paper's senior author. "We hope this tool, which displays zonation of carbohydrate and nitrogen metabolism, in addition to drug detoxification and alcohol degradation, will improve our ability to understand and predict the effects of toxins and new drugs on the liver."
More information: William J. McCarty et al, A Microfabricated Platform for Generating Physiologically-Relevant Hepatocyte Zonation, Scientific Reports (2016). DOI: 10.1038/srep26868