New technique takes guesswork out of IVF embryo selection

Researchers at the University of Adelaide have successfully trialed a new technique that could aid the process of choosing the "best" embryo for implantation, helping to boost the chances of pregnancy success from the very first IVF cycle.
The research—published today in the journal Molecular Reproduction and Development - has used highly advanced digital imaging techniques and mathematical modelling to show differences in the viability of embryos, which are not otherwise seen by the human eye under a microscope.
"It's fair to say that to date, strategies in IVF for picking the best embryo to transfer into the mother have been limited," says lead author Dr Hannah Brown, Postdoctoral Fellow with the University of Adelaide's Robinson Research Institute.
"There may be a number of embryos that look almost identical, and it's up to the embryologist to make a judgment call about which of them is best—that is, the most viable for a healthy pregnancy. That's a very difficult decision to make based on the little evidence available," she says.
"We know that many women who go through IVF aren't successful on the first cycle. This can be emotionally traumatizing and often becomes a very costly exercise depending on how many IVF cycles they go through.
"Using our knowledge of what is occurring in the biology of the embryo, we decided to see if there's more than meets the eye—the elements we can't see that distinguish the most healthy embryos, with the best developmental potential," Dr Brown says.
The key elements Dr Brown and her colleagues have considered are the quality of the embryo's metabolism and biomarkers for DNA damage that may have occurred during the embryo's in vitro development.
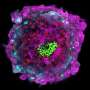
With assistance from researchers in the Centre for Nanoscale BioPhotonics (an Australian Research Centre of Excellence, also based at the University of Adelaide), the team has trialed a sophisticated, digital imaging technique—currently used for diagnosing cancer cells in patients—and mathematical modelling to create a "texture analysis" of the differences from one embryo to the next.
"These techniques provide a depth of analysis that is not otherwise discernible by the human eye. They're intentionally non-invasive to avoid causing any potential damage to the embryo or its environment," Dr Brown says.
"We have been successful on two fronts: in determining important differences between what would appear on the surface to be almost identical embryos, and in selecting those embryos that have had the best chance of a successful pregnancy.
"As we report in this new paper, these trials were conducted with mouse embryos. This is very promising work, and we are hopeful that in the years to come such a technique could be applied to IVF procedures.
"Our ultimate aim is to make the process of IVF more successful for couples, and to help produce the healthiest pregnancy possible for the benefit of the whole family," she says.
More information: Tiffany CY Tan et al, Grey level co-occurrence matrices (GLCM) to assess microstructural and textural changes in pre-implantation embryos, Molecular Reproduction and Development (2016). DOI: 10.1002/mrd.22680