Study unmasks the genetic complexity of cancer cells within the same tumor
A new study led by Cedars-Sinai investigators dramatically illustrates the complexity of cancer by identifying more than 2,000 genetic mutations in tissue samples of esophageal tumors. The findings reveal that even different areas of individual tumors have various genetic patterns.
The study results, published in the journal Nature Genetics, help explain why it is so difficult to battle cancer by targeting a specific genetic defect. A surgeon who performs a single biopsy on a patient's tumor can decode only part of the tumor and its genetic variations. Additionally, cancer cells constantly change their makeup.
"A tumor is not a single disease," said Dechen Lin, PhD, assistant professor and research scientist in the Division of Hematology and Oncology in the Cedars-Sinai Department of Medicine. "It's many diseases within the same person and over time. There are millions of cells in a tumor, and a significant proportion of them are different from each other." Lin was the project coordinator for the multicenter study.
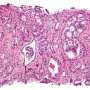
The cancer that the team studied, esophageal squamous cell carcinoma, is especially difficult to treat. The disease attacks the esophagus, the hollow tube that connects the throat to the stomach. The five-year survival rate for patients with esophageal cancer is about 20 percent, according to the American Cancer Society.
To create their catalog of mutations, the study's investigators called on high-powered computers to compile genetic data on 51 tumor samples taken from 13 patients. Through complex algorithms, they analyzed both the genes and the processes, known as epigenetics, that turned the genes' activities on and off within the cancer cells.
Using these techniques, the investigators identified 2,178 genetic variations in the sampled tumors. Dozens of the variations involved genes known to be associated with enabling the development of cancer. The most striking finding was that many important mutations were detected only in some areas of a tumor, highlighting the complexity of the cancer cells. This finding also demonstrated the potential for inaccurate interpretation of a cancer's genetic makeup using the single-biopsy method, which is the standard approach in the clinic.
Besides cataloging these genetic variations, the study's investigators reconstructed a "biography" of the tumors, showing when some of these variations first appeared in the life cycle of the disease.
"This study is on the leading edge of looking within a tumor for heterogeneity, or variations, across patients and within the same patient. It also is one of the very first studies to look at epigenetic changes from different areas within a single tumor in a global way," said Benjamin Berman, PhD, the study's co-senior author, an associate professor of Biomedical Sciences and co-director of the Cedars-Sinai Center for Bioinformatics and Functional Genomics.
To meet the challenge of integrating this diverse data, Huy Dinh, PhD, a project scientist in Berman's laboratory and one of the study's co-lead authors, developed innovative computational methods.
Looking ahead, the investigators plan to apply their analytic techniques to other cancers and explore the significance of the genetic and epigenetic changes that they have so far identified. They view their work as fundamental to developing effective, individualized therapies to combat the drug resistance that many cancer patients face during the course of their disease.
"Evidence suggests that tumor heterogeneity is one of the major causes of drug resistance and treatment failure in cancer," said H. Phillip Koeffler, MD, professor of Medicine and the Mark Goodson Chair in Oncology Research at Cedars-Sinai. "In light of this situation, deciphering the genomic diversity and evolution of tumors can provide a basis for identifying new targets and designing personalized medicine strategies." Koeffler was the other co-senior author of the study.
More information: Jia-Jie Hao et al. Spatial intratumoral heterogeneity and temporal clonal evolution in esophageal squamous cell carcinoma, Nature Genetics (2016). DOI: 10.1038/ng.3683