ISBM releases the updated version of Immune Response Template
Drug development has always been expensive and complicated. Many drug candidates are discontinued from clinical development during later stages, and it is not always possible to determine doses of medication or recruit a suitable group of patients for trials. ISBM offers a new approach to solve these problems: quantitative systems pharmacology (QSP) modeling. Diseases and biological systems are modeled in the most precise mechanistic way, and equations are based on data and basic knowledge. QSP models have the greatest planning horizon; applied during the early stages of research, they predict what the developers will observe during the second phase of clinical trials.
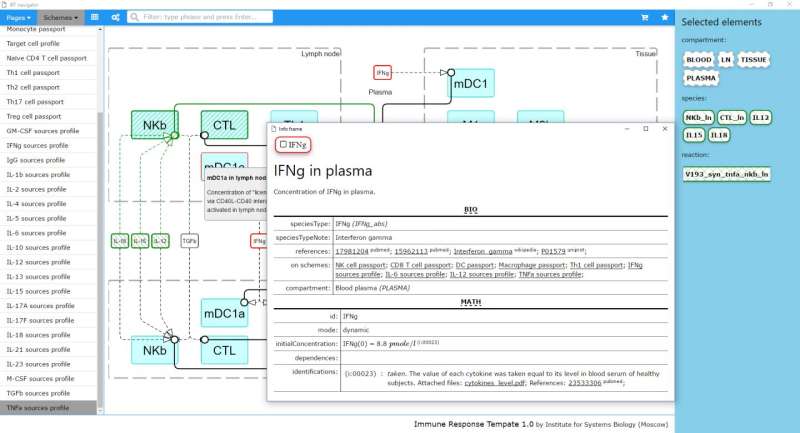
"It usually takes at least a half-year to build a QSP model describing the basics of immune response," said Oleg Demin Jr., a project leader. "IRT helps to shorten the time frame by designing a model template in few clicks. We describe the processes in the human body not only biologicaly, but also mathematically. Cells, cytokines and proteins are all the variables, and the immune system is represented as a complex system of equations. In fact, we have designed the human immune system on the computer and visualized it."
The software consists of an IRT database and the special IRT navigator to access it. The current database version describes about 10 types of immune cells, 10 surface molecules, 20 cytokines and 300 processes associated with immune response in human blood, lymph nodes and inflamed tissue. There are annotations for each process, cell and cytokine with cross references and links to the external databases in every interactive scheme. Platform users can see valuation parameters fitted with help of specific in vitro models, against in vitro data, or calculated using in vivo data measured for healthy humans and extended annotation of rate equations and parameters.
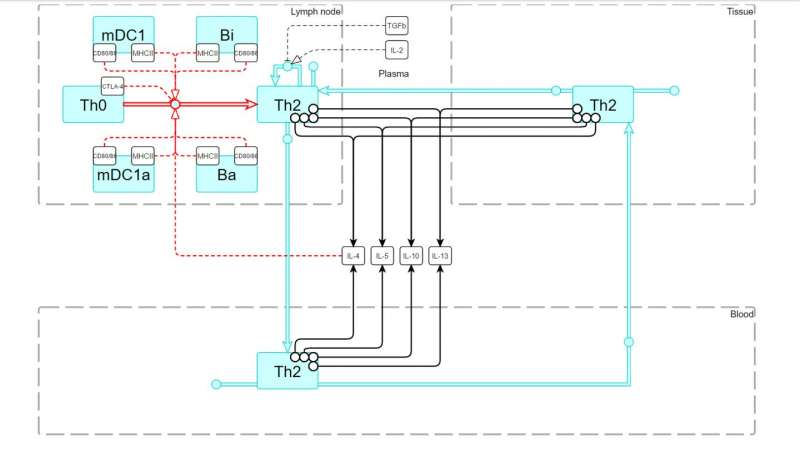
The user needs to choose the objects of interest, and the IRT navigator automatically generates and saves a model template with selected players, processes and corresponding parameters. Variation of these parameters simulates a virtual patient population which can be used for in silico trials. This approach improves the drug development process, especially in immune oncology field.