Spider web of cancer proteins reveals new drug possibilities

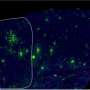
Scientists at Winship Cancer Institute of Emory University have mapped a vast spider web of interactions between proteins in lung cancer cells, as part of an effort to reach what was considered "undruggable."
This approach revealed new ways to target cells carrying mutations in cancer-causing genes. As an example, researchers showed sensitivity to an FDA-approved drug, palbociclib, for a gene that is commonly mutated in lung cancer cells, which is now being tested in a clinical study.
The results are published online in Nature Communications.
Many genes that drive the growth of cancer cells don't have any drugs available against them. For "tumor suppressor" genes, researchers are often not sure how to go after them. When the tumor suppressors are gone, cells often become more deranged, but there's no bullseye left to target. Exploiting the cancer cells' derangement remains a daunting challenge, says senior author Haian Fu, PhD.
"Our approach is to place tumor suppressors in the context of a network of cancer-associated proteins and link tumor suppressors to drugs through a known drug target protein," Fu says. "In this way, changes in a tumor suppressor may be linked with the response of the target to the connected drug."
The study is part of a push by the National Cancer Institute's Cancer Target Discovery and Development (CTD2) network to translate genomics data into therapeutic strategies, he says. Emory is a member of the NCI CTD2 network.
Fu holds the Winship Partner in Research endowed chair and is leader of Winship's Discovery and Developmental Therapeutics Program, director of the Emory Chemical Biology Discovery Center and professor of pharmacology and hematology and medical oncology. Co-corresponding author Fadlo Khuri, MD, maintains his professor appointment at Winship Cancer Institute and is now president of the American University of Beirut in Lebanon.
Cancer researchers have been searching for ways to target mutations in the gene STK11/LKB1, found in 15 to 25 percent of non-small cell lung cancers. The tumor suppressor STK11/LKB11 encodes an enzyme that is thought to regulate cell migration and metabolism.
One of the Winship team's newly identified interactions—a "thread" in the spider web—suggested that palbociclib, recently approved against metastatic breast cancer, may work against cells carrying mutations in LKB1, through LKB1's connection to CDK4, the target of palbociclib.
That prediction was supported by genomic data analysis and cell culture experiments: lung cancer cells with LKB1 defects showed a tendency of increased sensitivity to palbociclib. Now a study led by Taofeek Owonikoko, MD, at Winship is using LKB1 status as a biomarker for interpreting the effect of palbociclib.
How OncoPPI works
If cells are complex machines, then a number of ways exist for figuring out how the machines' parts, dominated by proteins, fit together. Some of them involve multiple washing steps to remove nonspecific partners after breaking cells apart, but FRET (Förster resonance energy transfer) does not. If two fluorescent molecules with colors that are near on the spectrum are close enough (less than 10 nanometers), that proximity can be detected by FRET.
Fu and his colleagues established a large-scale platform for tagging proteins with two different fluorescent molecules, introducing them into cancer cells, and then detecting interactions between the proteins. They call this network of cancer-associated proteins "OncoPPI."
Starting with a set of 83 lung cancer-related proteins, the team detected more than 260 interactions that were not known previously. They tested the interactions several times, in different orientations, and in other lung cancer cell lines with selected interactions to establish reliability. More than 80 percent of the interactions the researchers detected could be confirmed by another method (GST pulldown).
As an additional example to illustrate the utility of a protein interaction web, the team focused on the prominent oncoprotein Myc, which was also considered "undruggable." But the researchers could connect Myc indirectly through NSD3 to another protein called Brd4, against which inhibitors have been developed. Brd4 inhibitors are being currently tested in clinical trials. This finding revealed a new pathway Brd4-NSD3-Myc as potential targets for therapeutic intervention, Fu says.