Pathogen demonstrates genome flexibility in cystic fibrosis
Chronic lung infections can be devastating for patients with cystic fibrosis (CF), and infection by Burkholderia cenocepacia, one of the most common species found in cystic fibrosis patients, is often antibiotic resistant. In a study published today in Genome Research, scientists sequenced and phenotyped multiple B. cenocepacia isolates from 16 CF patients. They found extensive variation among isolates during chronic lung infection as well as changes in clinically relevant bacterial phenotypes.
"We expected, based on anecdotal observations and single strain reports, that the genome of B. cenocepacia was flexible, but we had no idea of the scope and scale of how promiscuous the gene content and genome architecture would be in a modest-sized patient cohort," said co-corresponding author, Corey Nislow, from University of British Columbia.
The researchers collected 215 isolates from 16 CF patients from the Canadian Burkholderia cepacia Complex Research and Referral Repository (CBCCRRR), with samples spanning a period of 2 to 20 years for each patient. Most patients demonstrated significantly decreased lung function during this time. Using whole genome sequencing, the genetic content of all isolates was profiled and genome assemblies were generated for 11 isolates. "By looking at changes in the genome over time, we were able to see patterns—common themes that help us to better understand how this particular species evolves in its environment and how CF patients become chronically infected," said study co-corresponding author Joshua Chang Mell, from Drexel University College of Medicine.
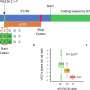
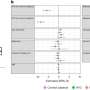
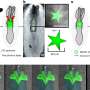
Similar to previous studies, the researchers found chromic infection of B. cenocepacia resulted in genome reduction, specifically loss of genes encoding non-essential functions, such as putative virulence genes. Phenotypic changes also occurred in a patient over time, including progressive decreases in motility and acute virulence, and changes in growth and biofilm formation. Although infections originated from a single strain, there was large phenotypic variation from samples taken later from the same patient at the same time, suggesting subsequent diversification within an infection.
While some isolates showed strong positive correlation between traits such as motility and biofilm formation, isolates from another patient showed an inverse correlation, suggesting the genetic architecture of the same trait may be distinct across strains.
Testing for associations between genetic variation and phenotypic differences, researchers identified numerous variants in genes associated with motility and biofilm formation. In addition, the loss of three genes previously associated with biofilm formation was correlated with both reduced motility and biofilm formation phenotypes in B. cenocepacia. The genetic determinants of motility and biofilm phenotypes may be promising targets for anti-virulence drugs.
"The outbreaks of B. cenocepacia in Canada and the UK in the 1990's have been largely contained by introduction of infection control measures, but we believe that, rather than 're-fighting the last war', the insights into which genotypic and phenotypic elements are pathogenic will let the B. cenocepacia community be proactive in responding to the next outbreak when it arrives," Nislow said.
More information: Lee AH-Y, Flibotte S, Sinha S, Paiero A, Ehrlich RL, Balashov S, Ehrlich GD, Zlosnik JEA, Mell JC, Nislow C. 2017. Phenotypic diversity and genotypic flexibility of Burkholderia cenocepacia during long-term chronic infection of cystic fibrosis lungs. Genome Res DOI: 10.1101/gr.213363.116