'Detective' iSHiRLoC investigates role of RNA, could help researchers study genesis of cancer
A team of University of Michigan scientists have developed a tool to help them study dysfunction in cells by tracking a molecule called microRNA, possibly giving them a way to determine how cancer occurs and spreads in the body.
In a person's cells, RNA receives instructions from DNA about how to build proteins, then directs the cell to build those proteins. The RNA that codes protein is called messenger RNA. However, scientists estimate that while upward of 95 percent of DNA transcribes into RNA, most of that RNA does not actually translate into proteins.
Some of these RNA strands break into smaller chunks called microRNAs, or miRNAs, which stop messenger RNA from producing proteins. In human cells, about 2,000 miRNAs regulate RNA function across the entire cell. Following the paths of miRNAs can tell researchers when and why RNAs in the cell become dysregulated, leading to disorders such as cancer.
"These RNAs that don't code protein, or are 'noncoding,' regulate everything that goes on in the cell, and when they become dysregulated—if they are in the wrong concentration or expressed at the wrong time—typically bad things, like cancer, happen," said Nils Walter, U-M professor of chemistry and biophysics.
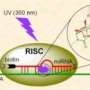
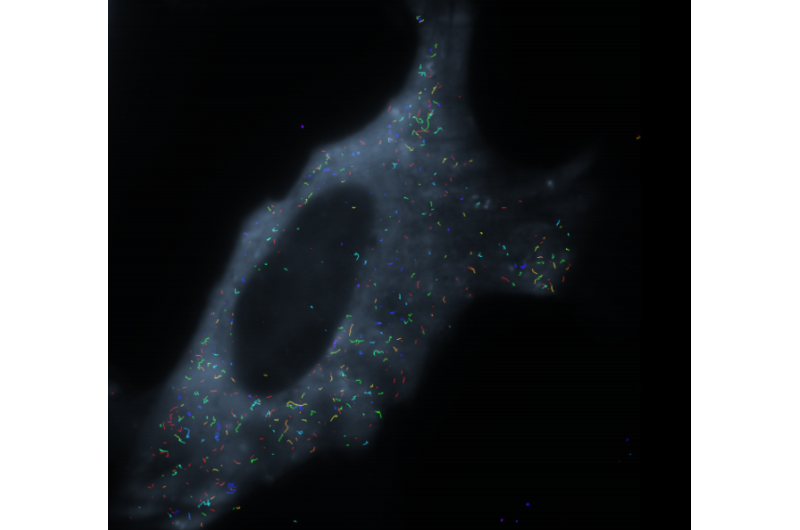
To track the miRNA molecules, the researchers attach what's called a fluorophore to the molecule. They excite the fluorophore, a dye molecule, with a laser light of a particular wavelength, causing the fluorophore to emit light of a different wavelength. Using a powerful microscope, the researchers can pick up the light, tracking the miRNA at a single-molecule level. They have called their tool iSHiRLoC, or "intracellular single molecule, high-resolution localization and counting." The study is published in the journal Cell Reports.
"In this paper, we've shown that we have the tools with which we can actually see single miRNAs in cells, and we can look at their movement from nucleus to cytoplasm and cytoplasm to nucleus," Walter said.
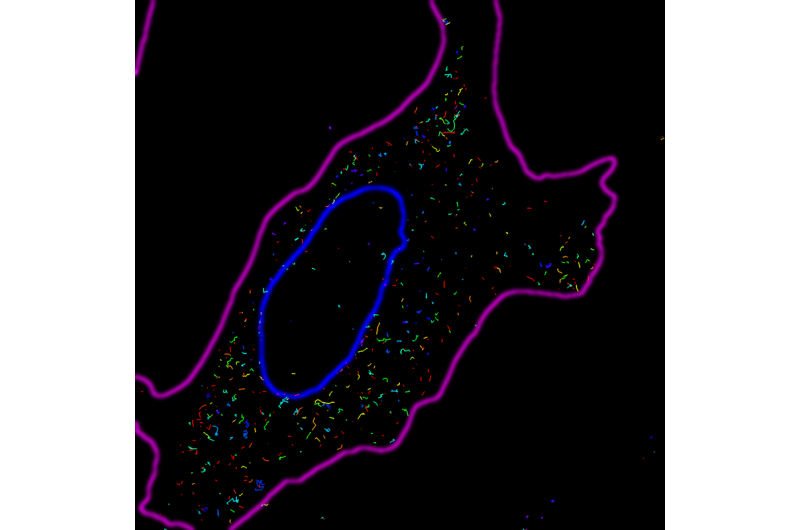
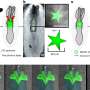
The microinjected miRNAs, which the researchers call "naked" miRNAs, have nothing bound to them, do not yet regulate and move through the cells too quickly for the researchers to track in real time. Fortunately, they slow down once they start regulating, so the researchers can follow their movements in two ways.
First, they observe what the miRNAs are doing in the living cell at different points in time after they've been injected a miRNA with the fluorophore attached. Second, they replicate the experiment in cells in which they freeze all motion at similar times through fixation after they inject the miRNAs to create snapshots of their nonregulating and regulating activity within the cell. Then, the researchers compare the microRNA activity in the live cell experiment with the snapshots from the fixed cells.
"By correlating the two numbers, we can figure out what fraction of the microRNAs are now engaged with the regulated messenger RNA target," Walter said. "Now we have a tool with which we can determine how quickly a microRNA engages a messenger RNA target, or how quickly it is lost from the cell and disappears entirely."
The researchers say that monitoring this activity can show them whether a drug targeted toward a particular cancer is working by showing whether the cell under attack is regulated or not.
Next, they plan to investigate a surprising discovery they made in the current study: the function of miRNAs in the nucleus. Previously, scientists thought miRNA only regulated RNA in the cytoplasm of the cell, but more evidence shows miRNA is present in the cell's nucleus where the DNA is housed. The U-M researchers want to study whether miRNAs are regulating the cell's nucleus, which would have an effect on the number of genes expressed through epigenetics.
"Or the miRNAs may just be sequestered away in the nucleus and not doing their jobs in the cytoplasm, which ultimately changes how much mRNA regulation they do," Walter said.
More information: Resolving Subcellular miRNA Trafficking and Turnover at Single-Molecule Resolution. DOI: dx.doi.org/10.1016/j.celrep.2017.03.075