Researchers home in on mutation profiles of clear cell endometrial cancer
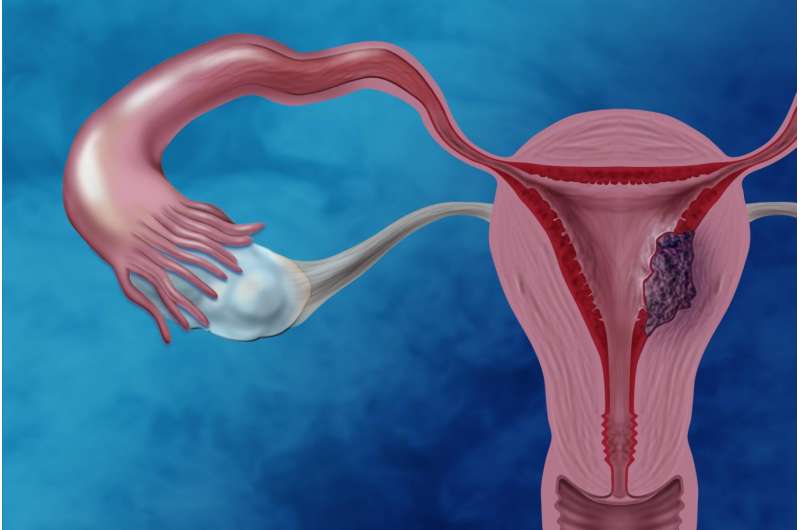
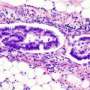
In the largest genomics study of clear cell endometrial cancer (CCEC) tumors to date, National Human Genome Research Institute (NHGRI) researchers and their collaborators have identified mutations in the TAF1 gene. They've also demonstrated that the mutation profiles of CCEC tumors are sometimes similar to other types of uterine cancer, namely serous endometrial cancers or endometrioid endometrial cancers. The study was published May 9, 2017, in the journal Cancer.
Endometrial cancer is the most commonly diagnosed gynecological cancer with more than 8,000 American women and 76,000 women worldwide dying from it every year. Fortunately, the most common type of endometrial cancer, endometrioid endometrial cancer, tends to diagnosed early and generally responds well to treatment; the five-year relative survival rate was reported as 91 percent in one study. However, women with CCEC or serous endometrial cancer, both rare types of endometrial cancer, generally have poorer clinical outcomes: One study reported that women with CCEC had a five-year relative survival rate of 65 percent, and women with serous endometrial cancer had a 45 percent five-year relative survival rate.
"Genomic mutations that contribute to clear cell endometrial cancer can vary from one patient to another," said Daphne W. Bell, Ph.D., senior author and senior investigator in NHGRI's Cancer Genetics and Comparative Genomics Branch. "Because the cancer is often diagnosed after it has spread beyond the uterus, the mortality rate is unacceptably high. We're trying to build a knowledge base to decipher the fundamental genomic changes that drive these cancers." Dr. Bell and her team specialize in basic research aimed at identifying genes that are mutated (changed) in the tumors of patients with clinically aggressive forms of endometrial cancer.
Dr. Bell and her team isolated DNA from tumor cells and from normal cells from patients with CCEC. Their goal was to identify genetic mutations in CCEC tumor cells that are absent from normal cells. The team sequenced the exome, the building blocks (nucleotides) in the protein-coding regions of the tumor and normal DNA from 16 CCEC patients, and found mutations in 222 genes in the tumors. The team narrowed this list of 222 genes to 22 "genes-of-interest" that represented genes mutated in more than one CCEC patient, genes previously implicated in CCEC, and/or genes that are mutated in clear cell ovarian cancer. The team further searched for mutations in these 22 genes-of-interest in tumor DNA from another 47 CCEC patients. Their results defined the frequency of mutation of the genes previously implicated in CCEC (TP53, PPP2R1A, PIK3R1, SPOP, PIK3CA, ARID1A and FBXW7) across the largest group of CCEC patients.
They also uncovered mutations in the TAF1 gene, a gene that had not been previously implicated in CCEC. TAF1 encodes part of a transcription factor complex, which regulates the expression of other genes throughout the genome. In addition, the researchers looked at patterns of gene mutations in tumors from CCEC patients and found that a substantial number of CCECs resembled serous endometrial cancer or endometrioid endometrial cancer.
The findings shed new light on the underlying genomic changes that are likely to be important in driving the development of CCEC, a rare but clinically aggressive form of endometrial cancer.