Researchers discover an important clue to help predict disease

The more information you have, the better able you are to predict what will happen next. Clinicians and health researchers often look at gene mutation to predict whether a fetus is at risk for a birth defect, or a person is at risk of developing a disease, but these predictions are not always accurate. University of Calgary researchers have discovered an important factor that changes our understanding of the relationship between gene mutations (genotype) and how they present in people (phenotype) that may, one day, help to improve this accuracy.
"The significance of this work is that it helps us understand how two individuals can have the same genetic mutation, but one can have a disease and the other one can be just fine," says Rebecca Green, Ph.D., a postdoctoral fellow in the Benedikt Hallgrimsson lab and first author of the study. "It happens commonly in cleft lip and palate; two kids will have the exact same mutation, but one will be born with cleft and one won't."
To better understand why, the researchers looked closely at gene expression levels. It has long been thought that changes in gene expression result in proportional changes in traits determined by those genes. Much of the variation in traits such as facial shape, height or blood pressure is determined by differences in levels of gene expression.
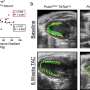
Using mouse embryos, they created a series of mutations to gradually decrease the expression level of a gene called Fgf8, which is important for proper development of the face. They started at 100 per cent and reduced it to 20 per cent of the typical level. Then they mapped the relationship between the two and found the relationship isn't proportional, there isn't a constant ratio, the relationship is non-linear.
"Our results showed even at 50 per cent of the normal level of gene expression you can be totally fine, but as it decreases further it doesn't mean the defect increases—you have a lot of different outcomes: from totally typical to essentially almost not even having a face," says Hallgrimsson, Ph.D., head of the Department of Cell Biology and Anatomy at the Cumming School of Medicine (CSM). "Those are the kinds of effects that result from a non-linear genotype/phenotype map. If you don't know what those maps look like you can't predict the phenotype from the genome."
The results are published in Nature Communications.
"This is a critical piece of understanding that's relevant to predicting disease from genotypes," says Hallgrimsson. "It adds to our knowledge of how genes can interact with each other and gives us one more answer for how variable outcomes can arise."
Hallgrimsson suspects if researchers looked at most really important genes they would have similar, highly non-linear genotype/phenotype map.
"The science of predicting phenotypes from genes is not as advanced as is commonly thought," says Hallgrimsson. "The prediction tools we've been using need to take non-linearity into account to refine and improve algorithms for using genomic data to predict variation. Those kinds of improvements will be critical for predicting disease risk."
This work also has significance for evolutionary biology. All organisms carry some amount of genetic variation. In order for a species to survive, it must be able to tolerate some amount of genetic variation and still develop and reproduce successfully. This study suggests that non-linearities in development are a major cause of this phenomenon.
More information: Rebecca M. Green et al. Developmental nonlinearity drives phenotypic robustness, Nature Communications (2017). DOI: 10.1038/s41467-017-02037-7