Looking beyond the 'magic bullet' approach to drug discovery

It's time to move beyond the traditional "magic bullet" approach for discovering new drugs and start leveraging the full complexity of Mother Nature, according to Vanderbilt Professor of Chemistry Brian Bachmann and Assistant Professor of Cell and Developmental Biology Jonathan Irish.
The researchers have developed a new process that can rapidly and inexpensively identify personalized cancer drugs derived from nature. They describe their new approach and report how it works on acute myeloid leukemia in a paper published Jan. 2, 2018, in the journal Nature Communications titled "Discovery of human cell selective effector molecules using single cell multiplexed activity metabolomics."
"The conventional 'magic bullet' approach to drug development is focused on identifying single molecules that kill disease organisms by disrupting single targets in single artificial cell lines. This has had some successes, but it also has a lot of limitations," said Bachmann, whose research focuses on searching for novel drugs produced by bacteria found in caves, and studying how these drugs are synthesized.
"Within the drug discovery community, there is a tendency to look for simple solutions," said Bachmann. "But life is very complex, as are many diseases, and we need to embrace that complexity to solve complex problems in human medicine."
Many diseases, including many cancers, consist of cells that possess multiple mutations and redundancies that allow them to compensate for a drug hitting a single target. And the artificial cell lines that researchers often use, like the HeLa cancer cells isolated from Henrietta Lacks in 1951, do not always behave and respond like naturally occurring—or "wild"—cancer cells found in a patient.
"We wanted to develop a technique that was able to simultaneously detect the effects of multiple compounds against multiple targets in multiple wild cancer cells mixed with normal cells, much as tumors occur in the body," said Irish, who studies the changes in human cells that lead to drug resistance.
In doing so, Bachmann and Irish have brought an old idea—searching for drug ingredients that occur abundantly in nature rather than developing them one at a time in a lab—into the 21st century with cutting-edge scientific tools and computational power.
The researchers began with a type of bacteria known for producing molecules that could affect human health. The bacteria produce thousands of these compounds, called secondary metabolites, to defend themselves from attacks or to attack their competitors. However, they are difficult to discover because the bacteria only produce them under very specific conditions, often in low abundance.
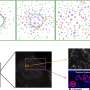
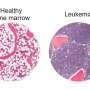
Using a pioneering technique developed in Bachmann's lab two years ago, the researchers stimulated the bacteria to produce large quantities of these chemical defenses. The researchers then combined samples of each individual metabolite with a mixture of healthy and diseased cells biopsied from real patients diagnosed with acute myeloid leukemia.
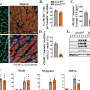
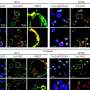
They then "bar-coded" the contents of each sample with fluorescent dyes and mixed the samples back together. The researchers used those fluorescent bar codes to quickly analyze how these thousands of metabolites affected the patients' biopsied cells, for better or worse, or not at all.
Advanced computational tools such as machine learning were used to process and recognize patterns in the enormous data sets that emerged from these thousands upon thousands of analyses.
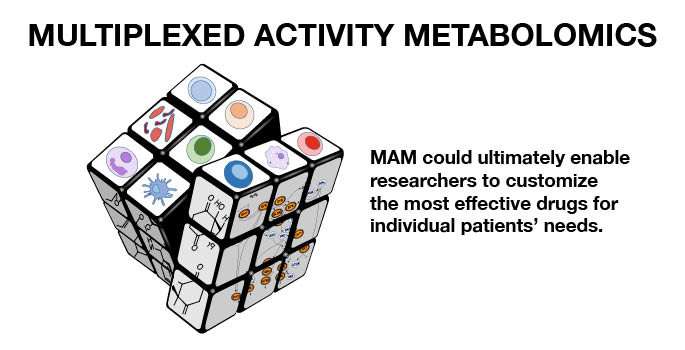
"We call this multiplexed activity metabolomics, or MAM, and it is potentially thousands of times less expensive than traditional methods. It also allows us to simultaneously ask many questions about what the compounds are doing to the cells, unlike traditional assays that each ask a single question," Bachmann said. "This is akin to a shotgun approach, casting a broad net across cellular responses, which can also detect the unexpected."
"This multiplexed approach contrasts with a targeted approach in that it performs a multitude of simultaneous tests and then sorts the results," Irish added. "It is left to us as researchers to identify the interesting results, such as metabolites that result specifically in death of one cell type, such as leukemia cells, and not healthy cells."
The net result is that MAM enables researchers to identify which metabolites or mixtures of metabolites target which type of diseased cells while sparing healthy cells, even down to even how well they might work in a particular patient.
Bachmann and Irish believe that by performing the discovery process with multiple patient samples representing the various major genetic types of cancer, molecules emerging from their pipeline could be developed for use in personalized therapeutics that are tailored to individual patients.
The importance of personalized forms of treatment were illustrated by the tests that the researchers performed on biopsy samples of two leukemia patients: a young woman with a treatable form of the cancer and an older adult male with a drug-resistant form of the disease. They determined that the same metabolites were more toxic to cancer cells in the treatable patient than those in the resistant patient. They also discovered metabolites that targeted the cancer cells in the resistant patient while doing relatively little harm to his healthy cells.
"Our new technology allows drug discovery using complex human tissues from patients, without having to experiment directly on people. By leapfrogging past artificial cell lines and animal models of disease, we think this will accelerate the discovery and approval of new therapeutics for complex human diseases such as cancer," Bachmann said.
The researchers don't plan to stop with cancer-killers. "In the future, we plan to use the system to look for more complex patterns, such as activating immune cells while suppressing cancer cells or differentiating a particular cell type into a useful type of specialized cell," Irish said. "Since we can look for molecules with positive and negative impacts at the same time, we imagine the MAM system being used for regenerative medicine as well as drug discovery."
More information: David C. Earl et al. Discovery of human cell selective effector molecules using single cell multiplexed activity metabolomics, Nature Communications (2017). DOI: 10.1038/s41467-017-02470-8