Real-time, web-based tool to revolutionise drug-resistant TB treatment
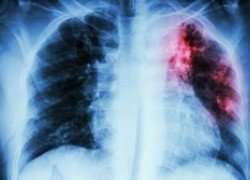
Melbourne researchers have designed a computer-generated model that will allow clinicians to tailor effective therapies for individual patients with multidrug-resistant tuberculosis (MDRTB), and as a result, reduce drug resistance globally.
The World Health Organization says TB is the leading cause of death globally from a single infectious agent – Mycobacterium tuberculosis – that typically affects the lungs, but can also infect other sites of the body. Multiple antibiotics are used to treat TB, however treatment is long, and the emergence of drug-resistant bacteria is a considerable threat to global health.
Researchers from The Peter Doherty Institute for Infection and Immunity (Doherty Institute) and the University of Melbourne's Bio21 Molecular Science and Biotechnology Institute (Bio21) used cutting-edge genome sequencing technology to identify an MDRTB mutation in a particular patient.
Armed with this information, University of Melbourne Ph.D. student Malancha Karmakar developed a three-dimensional (3-D) computational approach to determine that a drug being used to treat the patient called pyrazinamide was ineffective due to the mutation.
Rather than waiting two years (the standard treatment time for MDRTB) for the patient to complete treatment with pyrazinamide only to rule it redundant, they were able to replace it with a different drug immediately.
Study co-author and Medical Director of the Victorian Tuberculosis Program at the Doherty Institute, Associate Professor Justin Denholm, said the new tool was a game-changer. Whereas in the past it could take decades to identify the effectiveness of a TB drug, it could soon be a matter of hours.
"Right now, to use genetic tests to reliably predict whether a drug will be effective, someone needs to have been treated with a drug, failed that therapy and to be found to have a gene mutation," Associate Professor Denholm said.
"Then you need enough people around the world to have that pattern to inform what treatment you prescribe for that particular mutation. It can take years."
Co-author and University of Melbourne Professor David Ascher from Bio21 said by understanding how mutations work within 3-D space, the team could identify likely resistant mutations that have never arisen before.
"This is a huge boost to drug-resistant TB programs, which globally have to take a 'best guess' approach, where they give the same, standardised therapy to people with MDRTB, with little capacity to adjust that in real-time," Professor Ascher said.
Researchers say once the tool is fully developed it will be available through the web, meaning clinicians around the world could enter the resistance mutations into the system to determine effective treatment regimens for their patients.
This study was published in the American Journal of Respiratory and Critical Care Medicine.
More information: Malancha Karmakar et al. Analysis of a Novel pncA Mutation for Susceptibility to Pyrazinamide Therapy, American Journal of Respiratory and Critical Care Medicine (2018). DOI: 10.1164/rccm.201712-2572LE