AI accurately predicts effects of genetic mutations in biological dark matter
A new machine learning framework, dubbed ExPecto, can predict the effects of genetic mutations in the so-called "dark matter" regions of the human genome. ExPecto pinpoints how specific mutations can disrupt the way genes turn on and off throughout your body. Such disruptions in gene expression can sometimes have fatal consequences.
Using the method, its creators at the Flatiron Institute's Center for Computational Biology (CCB) in New York City and at Princeton University computed the genetic ramifications of more than 140 million mutations in different tissues. The researchers also precisely pinpointed mutations potentially responsible for increasing the risk of several immune-related diseases, including chronic hepatitis B virus (HBV) infection and Crohn's disease.
ExPecto could one day aid in the selection of drug therapies and help illuminate how evolution shaped our genetic code, the researchers report in a study published on July 16 in Nature Genetics.
"ExPecto can examine any genetic variant and predict its effect on gene expression," says principal investigator Olga Troyanskaya, deputy director of genomics at CCB and a professor at Princeton. "That's incredibly exciting."
Your DNA contains genes that serve as blueprints for building proteins, the workhorse molecules of our bodies responsible for carrying out important tasks such as ferrying oxygen, communicating with other cells and fighting infections. Protein-coding sequences make up less than two percent of your whole genome. All of these genes are present in cells throughout your body. This ubiquity means that protein-encoding genes vital to brain function, for instance, also exist in your digestive tract, lying dormant.
Genes are switched on and off by the other 98 percent of your genome, the "dark matter" portion that doesn't code for proteins. Most genetic mutations are found in this noncoding region. A mutation is essentially a genetic typo—an addition, deletion or alteration in the genomic sequence. Mutations in the noncoding region can sometimes cause genes to express or not express in the wrong part of your body at the wrong time, increasing the risk of diseases such as cancer.
Identifying the specific mutation responsible is difficult because the noncoding portion of DNA is so large. Previous studies compared the genomes of many individuals with a given disease, searching for mutations the individuals had in common. This approach, however, becomes increasingly tricky for rarer mutations. Furthermore, strings of DNA are sometimes inherited in large clusters, so scientists struggle to pinpoint which particular piece of genetic code is the troublemaker.
The study authors took a different approach. They developed ExPecto (named after the Patronus charm from the Harry Potter series) as a program that can read a raw sequence of DNA and predict the corresponding effect on gene expression.
ExPecto harnesses deep learning methods from artificial intelligence. Using a single reference genome, the researchers trained the program to understand how DNA controls gene expression across more than 200 different tissues and cell types. From this information, ExPecto can predict the effect of any mutation, even mutations that scientists have never seen before.
The researchers used ExPecto to predict the mutations that contribute to Crohn's disease, chronic HBV infection and Behçet's disease. Study co-author Chandra Theesfeld then experimentally verified the results. For all three diseases, she found that ExPecto's predicted candidate was a more promising potential contributor to the disease than those proposed by previous studies.
The researchers hope that ExPecto will one day help medical experts identify the genetic contributors to a patient's disease and develop therapies customized to the patient's genome. "Once you know which protein is affected and what the protein does, then you can design drugs that can fix the problem," says study co-author Jian Zhou, a Flatiron research fellow at CCB. For instance, "if you can't produce a certain protein, then you could design a therapy that makes up for the missing protein."
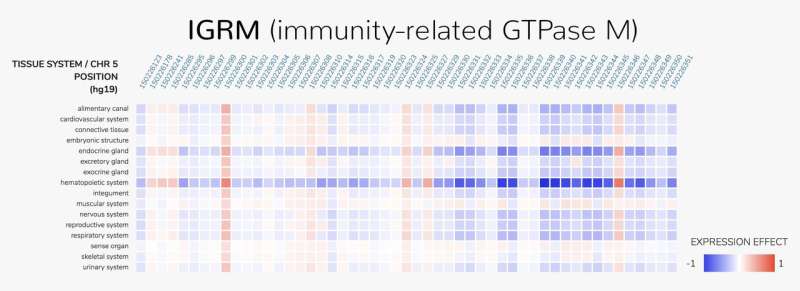
Anyone can access ExPecto's predictions of the effects of more than 140 million possible mutations near protein-encoding genes. These results are available online as part of HumanBase, a data-driven prediction system about human biology and disease developed by the research team. Visitors can type in a gene and see all the potential mutations that could affect that gene's expression in any of 218 tissues and cell types.
Zhou anticipates that ExPecto will be particularly insightful for studying the evolutionary consequences of mutations. He and his colleagues found, for instance, that mutations were less likely to affect genes expressed throughout the human body than genes specialized for one specific tissue type. "We don't have a full explanation yet," he says, but the result could be related to the robustness of more ubiquitous genes. An issue with a body-wide gene can have a higher likelihood of being fatal or otherwise preventing the individual from passing on his or her genetic information. "Evolution has already done the experiments for us," Zhou says.
More information: Jian Zhou et al, Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk, Nature Genetics (2018). DOI: 10.1038/s41588-018-0160-6