Researchers find first indicators of prognosis for the most aggressive breast cancer
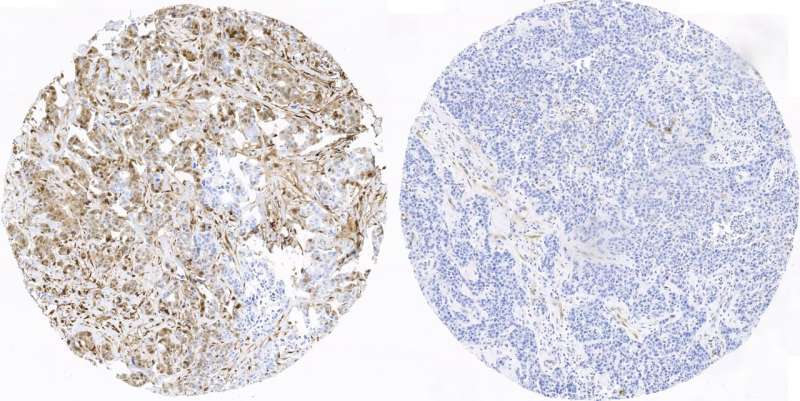
Triple negative breast cancer is the rarest, but also the most aggressive and hard to treat form of the disease. Researchers have so far been unable to identify markers that can classify patients by prognosis or probability of responding to different treatments. A group from the Spanish National Cancer Research Centre (CNIO) now reports a successful classification of triple breast cancer patients, which, for the first time, distinguishes those who can be cured from those who might suffer a relapse. It also identifies new pharmacological targets, and indicates that in patients with these targets, combined treatments with existing drugs could be effective. Their study is published in Nature Communications.
Specifically, they have identified six protein kinases whose functional status predicts the evolution of triple negative breast cancer. Furthermore, they found a way for these proteins to be studied in hospitals, so that in the future, it might be a regular clinical test, just as genetic profile analysis of any tumour is today.
"Until now, it has not been possible to establish a link between the presence of certain mutations in triple negative breast cancer and a prognosis or response to drug treatment," explained oncologist Miguel Ángel Quintela, director of the Breast Cancer Clinical Research Unit at CNIO and lead author on the research paper that is being published now. "We have shown for the first time that proteomics can be used to predict the evolution of triple negative breast cancer, and to select combinations of pairs of drugs as candidates for clinical trials."
From genomics to proteomics
Decades of research into cancer genomics have identified dominant gene mutations in many cancers that determine the progression of the tumour and guide the design of personalised treatment pathways. These therapies targeting the specific biology of each tumour, more effective than conventional chemotherapy, are largely responsible for the advances made in recent years in cancer treatments.
But triple negative breast cancer is caused by numerous mutations, which act in conjunction and in unique combinations for each patient. So far, it has not been possible to find dominant gene mutations that provide an indicator of prognosis or response to drug treatment.
As the researchers explain in Nature Communications, "Although studies based on genetics have provided unprecedented knowledge about breast cancer and its subtypes, in triple negative breast cancer, this new information has revealed major heterogeneity that has prevented us, so far, from defining prognostic or predictive factors. [This fact] has meant that conventional chemotherapy is still the main therapeutic option in triple negative breast cancer."
For that reason, "it would be extremely interesting" for this type of cancer to have a classification system associated with different therapeutic options, write the authors.
With this aim, CNIO researchers chose not to analyse the genes involved in triple negative breast cancer, but rather their product: the proteins whose synthesis is ordered by those genes. Their hypothesis was that the numerous genetic alterations of patients could translate into recognisable patterns of the functional status of all the tumour's proteins—this is the proteome.
The researchers were successful. In samples of tumours from 34 patients, they found more than 2 million biochemical markers of the activation of tumour proteins, and with the help of sophisticated bioinformatic tools, they determined that among all these signals, there is a precise combination found only in patients who relapse. These proteins are activated through kinases—which are in turn proteins—and so the next step was to find the kinases responsible for that specific pattern. Finally, the analysis identified the six kinases responsible for the activation pattern characteristic of the proteome of patients who relapse.
Six kinases indicate probable relapse
Some of these kinases had been studied previously. "But until now, there was no reason to home in on them," explained Quintela.
The validation of their findings with 170 patients confirmed the value of these six kinases as a marker. Patients in whom none of these proteins was activate had a 95 percent chance of being cured, or at least not suffering a relapse 12 years after treatment. However, if even one of the six kinases was active, the risk of relapse multiplied by ten.
These six kinases can be inhibited using drugs, and there are already drugs in use against some of them. Furthermore, to prove the clinical relevance of their findings, the researchers studied xenografts from human-derived tumours transplanted into mice. They determined anti-tumour activity of 15 different combinations of drugs, and related them with the activation profile of the six kinases.
They found combinations that achieved "promising anti-tumour activity," according to the Nature Communications article. Specifically, 15 combinations in 10 different models—150 situations—achieved a superior therapeutic effect to the sum total of the therapeutic effects of each drug separately in 99.3 percent of cases.
Toward a new clinical trial for kinase activation
Analysis of the functional status of proteins cannot currently be conducted as a routine test in hospitals, but the authors have translated the activation patterns of the kinases into indicators of immunohistochemistry, which can be analysed easily in hospitals. The objective is for the study of the six kinases identified to become a regular clinical test in the future, similar to the genetic profiling of any tumour is today.
Currently, researchers are focusing on "validating these markers in other stages of diseases, standardising kinase determinations by way of a diagnostic test, and organising clinical trials using the therapeutic combinations described in this paper on patients with advanced disease," said Quintela.
More information: Ivana Zagorac et al, In vivo phosphoproteomics reveals kinase activity profiles that predict treatment outcome in triple-negative breast cancer, Nature Communications (2018). DOI: 10.1038/s41467-018-05742-z