New information to understand regulation of muscle function in muscular dystrophy patients
Scientists from the University of Jyväskylä, Finland, and from University of Cambridge, United Kingdom, have found new functions and structural details of muscle attachment structures. The research is important for the basic understanding of regulation of muscle function, for instance, in muscular dystrophy patients and for muscle adaptation in physical activity and sports.
In multicellular organisms, cell adhesion to other cells and to the extracellular matrix is very tightly regulated. Depending on the tissue, the attachment structures are adjusted to tolerate various pulling forces, and sometimes, the attachments are opened and reorganized. For force tolerance, adhesion proteins in the cellular membrane need to be connected to the cytoskeleton, and this connection is the key regulatory point. For several years, the research groups of Professor Jari Ylänne at the University of Jyväskylä and Professor Nick H. Brown at the University of Cambridge have studied so-called mechanosensor proteins, whose functions change according to pulling force. The research teams found several new mechanisms regarding how the structure and function of muscle attachment sites are regulated during development.
Novel functions for proteins
Researchers utilized the flight muscles of Drosophila melanogaster fruit flies. These are the muscles in flies that most closely resemble human skeletal muscles, and their attachment sites are similar to myotendinous junctions, the connections between muscle and tendons. Drosophila flies are a very useful model organism—their development can be relatively easily followed under microscope, thousands of mutant and fluorescence protein marker fly lines are available, and genetic modification and genetic crossing are relative easy. Furthermore, flies often have fewer genes affecting the same functions in humans, and thus the effects of individual mutations are easier to observe.
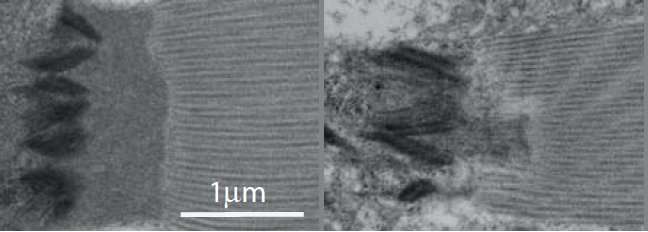

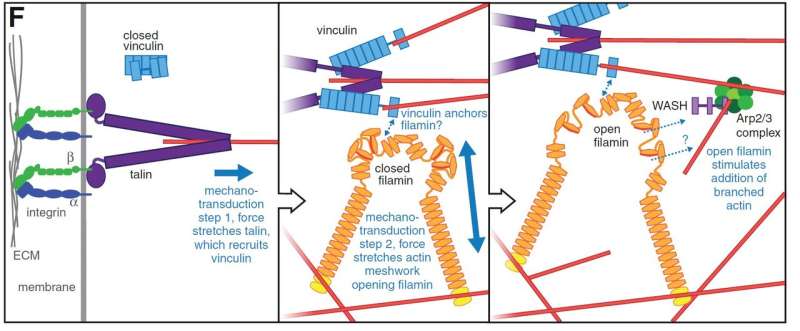
"By combining efforts between the two groups, we found a new functional link between three mechanosensor proteins. These proteins act in concert to strengthen the attachment sites when the muscle contracts. Alterations of these proteins or their specific mechanosensory sites caused disorganization of muscle attachment sites and reduced the flying capacity of the flies," says Professor Jari Ylänne from Department of Biological and Environmental Science and Nanoscience Center at the University of Jyväskylä.
More information: Hannah J Green et al. Novel functions for integrin-associated proteins revealed by analysis of myofibril attachment in Drosophila, eLife (2018). DOI: 10.7554/eLife.35783