Insulin tracked through cell using new method combining database, experimental results

High or low concentrations of insulin activate different cell signaling pathways, according to a new scientific method that combines data from multiple databases and large-scale lab experiments. This ongoing research project may help unveil better approaches to understand the causes of and potential therapies for type 2 diabetes.
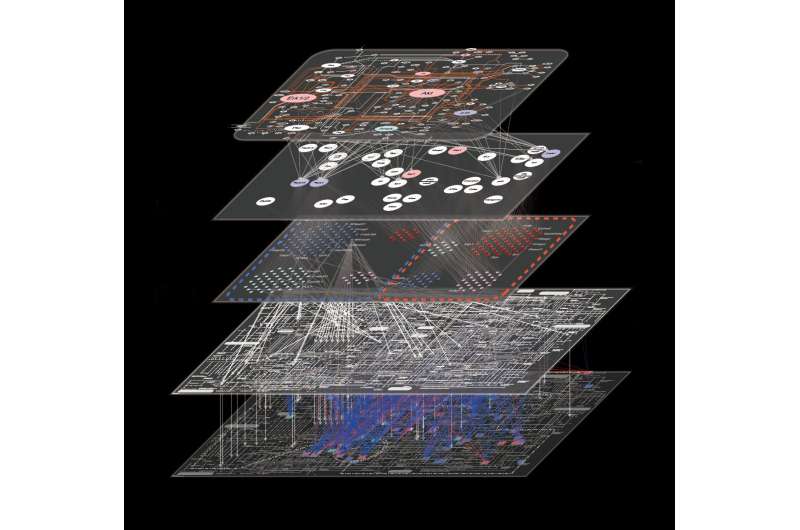
Scientists already have methods to understand all the genes (genomics), proteins (proteomics), or metabolites (metabolomics) within a cell, but not all of these different types of molecules at the same time. University of Tokyo researchers are pioneering the new trans-omics approach that combines all of these previously individual fields—the different "omics—to understand the interactions between molecules inside cells in a comprehensive, highly detailed way.
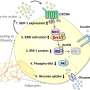
"Our results look almost like a subway map. Each molecule that insulin influences, directly or indirectly, is like a station. But a map is not very useful if you do not know the route. Our method combines database information with new experimental data to show how the different stations, or molecules, connect after receiving the insulin signal," said Professor Shinya Kuroda.
Professor Shinya Kuroda has been pioneering trans-omics approaches since the field started to emerge around 2013. Project Researcher Kentaro Kawata is the first author of the recent paper and completed the research as part of his doctoral studies.
Kuroda and his team are interested in how cells send signals to control their internal processes, especially how those signals change throughout the day. Insulin is a highly dynamic molecule and the hormone that helps control how the body turns sugar from food into energy. Improper regulation of insulin leads to diabetes.
"Type 2 diabetes is a complex disease, but it can be simply understood as an impairment of the temporal pattern of the body's response to insulin," said Kawata.
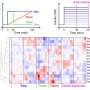
The research team uncovered that high and low concentrations of insulin activate different genes and metabolic processes. Cells respond not just to the presence or absence of insulin, but can interpret and respond differently to varying concentrations of insulin to control distinct biological processes.
The research team is currently conducting experiments to verify the results they find from their trans-omic data analysis are confirmed by studies with live mice. Detailed understanding of the dynamics of how and when insulin activates so many other molecules within cells may make it possible to design more effective insulin regimens for people with diabetes.
"Once we have mapped the large-scale network, we can identify potential drug targets," said Kuroda.
More information: Kawata K, Hatano A,i Yugi K, Kubota H, Sano T, Fujii M, Tomizawa Y, Kokaji T, Tanaka KY, Uda S, Suzuki Y, Matsumoto M, Nakayama KI, Kaori Saitoh K, Kato K, Ueno A, Ohishi M, Hirayama A, Soga T, and Kuroda S. 11 September 2018. Trans-omic analysis reveals selective responses to induced and basal insulin across signaling, transcriptional, and metabolic networks. iScience. 10.1016/j.isci.2018.07.022