The role of PRMT1-mediated alternative splicing in dilated cardiomyopathy
In a study published in iScience, Professor Akiyoshi Fukamizu of the University of Tsukuba has reported the discovery of the important role of PRMT1 in dilated cardiomyopathy (DCM).
The heart pumps blood to all organs and tissues. In the United States, the mortality rate due to cardiomyopathies is greater than 10,000 deaths per year. On the other hand, in one of the most common types of heart disease, DCM, cardiac muscle enlarges and weakens, preventing the heart from pumping blood efficiently. Moreover, at the time of listing for heart transplantation, the median age is four years, and over 10 percent of children affected with DCM have died at age two. While it is known that mutations in the alternative splicing regulator RBM20 gene have been shown to cause human DCM, genetic information such as gene expression and alternative splicing related to DCMs remains unidentified.
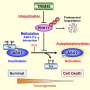
Arginine methylation is a posttranslational modification of proteins, and is catalyzed by protein arginine methyltransferases (PRMTs). PRMT1, amajormethyltransferase active in mammalian cells, regulates cellular functions such as gene expression, DNA replication, and cell proliferation. It is of note that these functions were identified by experiments with culture cells, while physiological roles of arginine methylation have been largely unknown.
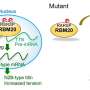
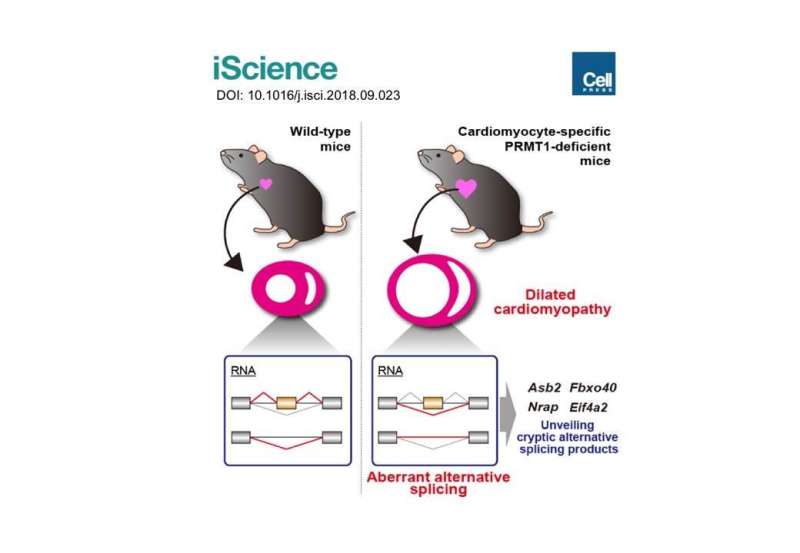
Previous studies showed the expression levels of PRMT1 were changed in the heart tissue of heart failure patients. However, we did not understand the role of PRMT1 in the normal or the failing heart. To address this question, researchers generated heart-specific PRMT1 knockout mice, namely PRMT1-cKO mice, which lack PRMT1 in cardiomyocytes, the heart muscles. Interestingly, the PRMT1-cKO mice developed severe DCM from the juvenile stage and sudden death, suggesting that PRMT1 is critical for proper cardiac function.
To uncover the mechanism underlying heart dysfunction in PRMT1-cKO mice, the researchers focused on messenger RNA (mRNA) alternative splicing, because some splicing regulators were known as methylation targets of PRMT1. Alternative splicing is a cellular system that produces a variety of proteins from a genome by selecting which part of the genes should be translated to proteins. Using high-throughput RNA sequencing (RNA-seq) for gene analysis, the researchers found aberrant mRNA splicing patterns of multiple genes in the PRMT1-deficient heart, suggesting that dysregulation of alternative splicing is strongly linked to the defective cardiac function observed in PRMT1-cKO mice.
Recently, a number of studies have suggested that abnormal mRNA splicing is related to the pathogenesis of DCM; however, its regulatory mechanisms are largely unknown. "Our findings may become a breakthrough for understanding the mechanism of DCM," says Akiyoshi Fukamizu. "DCM has serious prognosis, especially in cases of patients at young ages, including infants. We hope our study will benefit the development of new treatments for DCM patients."
More information: Kazuya Murata et al. PRMT1 Deficiency in Mouse Juvenile Heart Induces Dilated Cardiomyopathy and Reveals Cryptic Alternative Splicing Products, iScience (2018). DOI: 10.1016/j.isci.2018.09.023