Study reveals key details about bacterium that increases risk for stomach cancer
More than half of the people in the world host colonies of a bacterium called Helicobacter pylori in their stomachs.
Although it's harmless to many, H. pylori can cause stomach cancer as well as ulcers and other gastric conditions. Doctors tend to prescribe multiple antibiotics to defeat the microbe, but that strategy can lead to antibiotic-resistant superbugs.
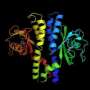
Now, a finding by UCLA scientists may lead to a better approach. The researchers have determined the molecular structure of a protein that enables H. pylori to stay alive in the stomach, and elucidated the mechanism by which that protein works.
Z. Hong Zhou, the study's corresponding author and a UCLA professor of microbiology, immunology and molecular genetics, said the findings answer questions that have been sought ever since 2005, when two Australian scientists won a Nobel Prize for their discovery of H. pylori and its role in gastritis and peptic ulcer disease.
The UCLA study, which was published online by Science Advances, was co-led by Keith Munson, a recently retired senior researcher in UCLA's Division of Digestive Diseases.
H. pylori thrives in the harsh environment of the stomach due to its urea channel, a protein in the bacterium's inner cell membrane that detects the environment's acidity and acts as a gate. When conditions in the stomach grow too acidic, the urea channel opens to let in a compound called urea. Urea is normally excreted as a waste product in urine, but it also can be found in relatively small concentrations in the stomach. H. pylori uses it as raw material for neutralizing the acid that otherwise would kill the bacterium.
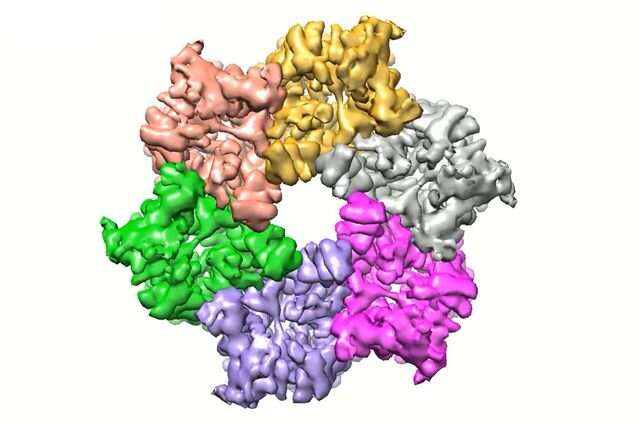
The research revealed the three-dimensional molecular structure of the urea channel, both when it is open and when it's closed, using cryo-electron microscopy, or cryo-EM, an imaging technique that detects electrons rebounding from frozen samples. Comparing the open and closed channels offered insight into the changes that take place when the "gate" opens, and the cryo-EM images provided previously unseen details that are important for understanding the activity of the protein, which is shaped like a hexagonal prism.
Zhou, Munson and their colleagues also engineered variations of the urea channel, shuffling different amino acids into key spots. They tested those variations at different levels of acidity to see which substitutions interfered with the protein's action. The experiments enabled the scientists to identify which parts of the urea channel are involved in sensing acidity and altering its shape to let in urea.
The UCLA discovery could lead to future research on ways to fight H. pylori more effectively by sabotaging its survival mechanism.
"The urea channel is a viable drug target for eradication of this human pathogen, which remains a significant health risk throughout the world," said Zhou, who also is director of the UCLA Electron Imaging Center for NanoMachines at the California NanoSystems Institute at UCLA.
The findings also represent a technical advance.
"We believe this is the highest resolution yet attained by cryo-EM for a membrane protein showing minimal mass outside the membrane," Zhou said.
More information: Yanxiang Cui et al. pH-dependent gating mechanism of the Helicobacter pylori urea channel revealed by cryo-EM, Science Advances (2019). DOI: 10.1126/sciadv.aav8423