Multi-modal analysis platform for capturing tumor-derived extracellular vesicles

Extracellular vesicles are arguably the next big thing in cancer diagnostics. They are membrane-bound particles shed by any cell in the body, and involved in intercellular communication. Analyzing these biomarkers seems worthwhile, as they contain a tremendous amount of information on their tissue of origin. Researchers at the University of Twente, in collaboration with colleagues from Wageningen University, are working on a multi-modal analysis platform for the specific capture of tumor-derived extracellular vesicles (tdEVs).
Minimally invasive diagnosis
It has been shown that the concentration of tumor-derived extracellular vesicles (tdEVs) in the blood of metastatic cancer patients correlates with their probability of survival. Simply by counting tdEVs in blood, patients can be monitored and treatment strategy adapted. This approach presents a significant advantage over conventional imaging techniques like MRI, which can only be employed a few times per year and cannot detect small tumors. In contrast, for EV analysis, only a blood sample is required, and such a liquid biopsy is much less invasive.
Still, before tdEVs can reliably be used in the clinic, they need to be more closely studied, which is a significant challenge due to: (1) their very small size, and (2) the vast number of other particles in blood with a very similar size. In other words, highly sensitive and specific methods are needed to detect tdEVs.
Multi-parametric characterization
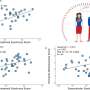
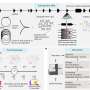
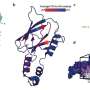
A collaboration between the UT's Medical Cell BioPhysics group (MCBP) and Applied Microfluidics for BioEngineering Research group (AMBER), together with Wageningen University, has led to the development of a platform for the reliable isolation of tdEVs followed by their individual multi-parametric characterization using scanning electron microscopy (SEM), Raman spectroscopy and atomic force microscopy (AFM) to acquire comprehensive information on the isolated particles. On a chemically modified substrate, tdEVs were specifically captured, which was confirmed by SEM. AFM yielded higher-resolution, 3-D information about their morphology.
Most notably, the chemical composition of single particles was disclosed by Raman spectroscopy. Aided by machine learning, the team was able to use this chemical inspection technique to identify a chemical fingerprint unique to tdEVs. Studying the particles individually, the team was able to verify the origin of the biomarkers, which was confirmed by combining the results obtained from the various instruments. Because if it looks like a tdEV, feels like a tdEV and smells like a tdEV, it must be a tdEV.
More information: Pepijn Beekman et al. Immuno-capture of extracellular vesicles for individual multi-modal characterization using AFM, SEM and Raman spectroscopy, Lab on a Chip (2019). DOI: 10.1039/c9lc00081j