Deep inside the brain: Mapping the dense neural networks in the cerebral cortex

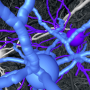
Mammalian brains, with their unmatched number of nerve cells and density of communication, are the most complex networks known. While methods to analyze neuronal networks sparsely have been available for decades, the dense mapping of neuronal circuits is a major scientific challenge. Researchers from the MPI for Brain Research have now conducted connectomic mapping of brain tissue from the cerebral cortex and quantified the possible imprint of learning in the circuit.
Brains contain extremely densely packed networks of membranous connections used by about 86 billion nerve cells for communication between one other. Since each nerve cell in the cerebral cortex of mammalian brains communicates with about 1,000 other nerve cells via synapses placed along these connections over long distances, there are about 5 million kilometers of neural wiring packed into our skulls—more than 10 times longer than all highways on Earth. The cables in mammalian brains are as thin as 50 to 100 nanometers in diameter; the resulting cable convolute is of such density and magnitude that for more than 100 years, researchers have been able to map only a miniscule fraction of neurons in a given piece of brain.
Currently, thanks to the development of faster electron microscopic techniques and more efficient image analysis routines, the dense mapping of neuronal networks has become possible. The novel field of connectomics has been pursuing the dense mapping of ever larger circuits in several species and brain regions.
In the work published today in Science, Max Planck Director Moritz Helmstaedter and his team imaged and analyzed a piece of tissue from the cerebral cortex of a four-week-old mouse obtained via biopsy from the somatosensory cortex, a part of the cortex occupied with the representation and processing of touch. Here, the researchers applied optimized AI-based image processing and efficient human-machine interaction to analyze about 400,000 synapses and about 2.7 meters of neuronal cable in the volume. They produced a connectome between about 7,000 axons and about 3,700 postsynaptic neurites, yielding a connectome about 26 times larger than the one obtained from the mouse retina more than five years ago. Importantly, this reconstruction was larger and about 33 times more efficient than the connectomic mapping of the retina, setting a new benchmark for dense connectomic reconstruction in the mammalian brain.
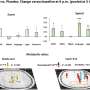
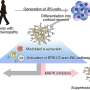
Fueled by this this methodological breakthrough in connectomics, the researchers analyzed the connectome for the patterns of circuitry. In particular, they asked what fraction of the circuit showed properties that were consistent with the growth of synapses, mechanisms known to contribute to circuit formation and learning. Alessandro Motta, first author of the study and an electrical engineer by training, used particular configurations of synapse pairs to study the degree to which they were in agreement with activity-related learning processes ("LTP").
"Because some models of synaptic plasticity make concrete predictions about the increase in synaptic weight when learning to, say, identify a tree or a cat, we were able to extract the imprint of such potential processes even from a static snapshot of the circuit," explains Motta. Since the mouse had lived a normal laboratory life until the brain biopsy at four weeks of age, the scientists argue that the degree to which circuits are shaped by learning in "normal" sensory states can be mapped using their approach.
"We were surprised how much information and precision is found even in a still relatively small piece of cortex," says Helmstaedter, "The extraction of the likely learned circuit fraction was an especially major eye opener for us."
The reported methods may have substantial implications for the transfer of insights about biological intelligence to artificial intelligence. "The goal of mapping neuronal networks in the cerebral cortex is a major scientific adventure, also because we hope to be able to extract information about how the brain is such an efficient computer, unlike today's AI," says Helmstaedter.
He describes a research field with major players including Google and the Intelligence Advanced Research Projects Activity (IARPA) in the U.S. "The ambition to learn from biological neuronal networks about the future of artificial neuronal networks is shared by major initiatives worldwide. We are very proud about having achieved the first milestone, a dense local cortical connectome, using exclusively public funding from the Max Planck Society."
After almost a decade of work, the researchers are enthusiastic about their achievements. "Being able to take a piece of cortex, process it diligently, and then obtain the entire communication map from that beautiful network is what we have been working for over the last decade," says Helmstaedter.
The researchers conclude: "We think that our methods, applied over a large range of cortical tissues from different brain areas, cortical layers, developmental time points, and species will tell us how evolution has designed these networks, and what impact experience has on shaping their fine-grained structure. Moreover, connectomic screening will allow the description of circuit phenotypes of psychiatric and related disorders—and tell us to what degree some important brain disorders are, in fact, connectopathies, circuit diseases."
More information: "Dense connectomic reconstruction in layer 4 of the somatosensory cortex" Science (2019). science.sciencemag.org/lookup/ … 1126/science.aay3134