Scientist uses 'mini brains' to model how to prevent development of abnormally small heads
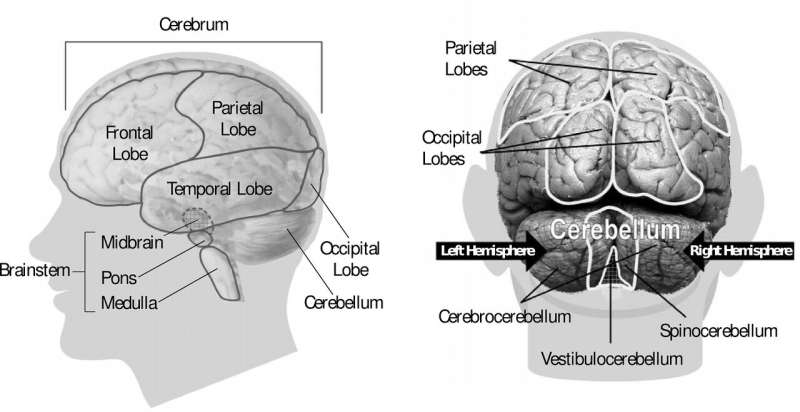
A City of Hope scientist is one step closer to discovering what weakens a pathogen that appears to cause babies to be born with abnormally small heads.
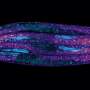
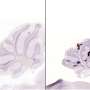
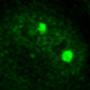
Interestingly, it takes studying "mini brains" to understand why certain unborn babies infected with cytomegalovirus (CMV) enter the world with shrunken brains, said Yanhong Shi, Ph.D., senior author of the new study, director of the Division of Stem Cell Biology Research and the Herbert Horvitz Professor in Neuroscience at City of Hope.
In the United States, the most common cause of infectious-related birth defects is CMV. About 1 in 5 babies with congenital CMV infection will have birth defects or other long-term health problems. Among those congenital conditions is microcephaly, or abnormally small heads, a concern many soon-to-be mothers had during the 2015 Zika outbreak. However, CMV is a far more common culprit for microcephaly, Shi said.
"We are among the first to model human CMV-induced microcephaly using brain organoids. This is a first step to one day studying more complex neurological complications such as Alzheimer's disease and Parkinson's disease," Shi said, adding that much more developed brain organoids, as the mini brains are more popularly known, are needed for scientists to study complex nervous system diseases that develop later in life.
The study, published in Cell Reports Medicine on March 25, solves a problem that has befuddled scientists for decades—how to create an experimental model that can mimic the complexities of the human brain in order to study neurological disorders. Until recently, scientists were restrained to studying the problem mostly in two dimensional models in a petri dish because they couldn't replicate many key features of neurological disorders in animal models. Notably, animals cannot be used to study human CMV (HCMV)-specific brain disorders because the disease is specific to humans.
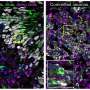
Shi and her colleagues, however, found that a strain of HCMV called TB40/E appeared to replicate what HCMV does to an unborn baby's brain in the transition between the first and second trimester. The TB40/E-infected brain organoids were significantly smaller than the control models. Of the 10 genes that were reduced, three were related to calcium signaling, an indication that brain connections were not being made and that the brain's electrical network was not functioning properly. Further testing showed that TB40/E affected critical genes involved in brain development, including ones responsible for the development of the hippocampus, the center of learning and memory.
"A similar organoid strategy can be used to understand how infection by the SARS-CoV-2 virus leads to COVID-19 so that we can test potential therapies for the disease," said Guoqiang Sun, Ph.D., lead author of the study and a staff scientist in the Department of Developmental and Stem Cell Biology at Beckman Research Institute of City of Hope.
To take the study one step further, Shi and her colleagues collaborated across disciplines with Don Diamond, Ph.D., professor in the Department of Hematology & Hematopoietic Cell Transplantation at City of Hope. Diamond has been studying CMV for three decades and is developing vaccines to prevent congenital CMV infection.
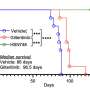
The City of Hope scientists tested what could one day prevent or lessen the birth defects created by HCMV-induced microcephaly. They introduced a protective immune system antibody currently in development in the Diamond Lab. When tested in the brain organoid model, it appeared early intervention with these "neutralizing antibodies" may prevent or reduce the most severe consequences of HCMV infection.
"Now that we have a model that replicates how HCMV-induced microcephaly happens, we can use it to test antiviral agents," Shi said. "We can now start looking for real-world solutions."
More information: Guoqiang Sun et al, Modeling Human Cytomegalovirus-Induced Microcephaly in Human iPSC-Derived Brain Organoids, Cell Reports Medicine (2020). DOI: 10.1016/j.xcrm.2020.100002