Allergic asthma transforms cells of the airway
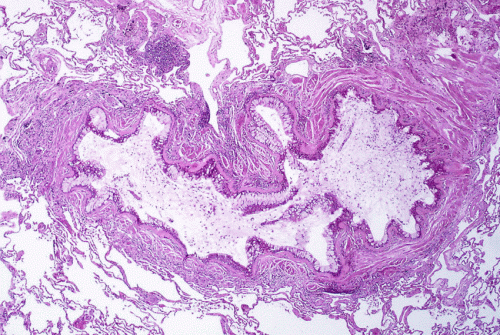
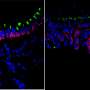
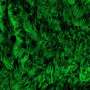
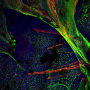
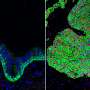
New research at National Jewish Health shows that allergic asthma fundamentally transforms the human airway, reducing cells' ability to remove pollutants and fight infections. Allergic asthma, the most common form of the disease, is driven by the signaling molecule IL-13. When researchers added IL-13 to cells extracted from human airways, it weakened the cells' innate immune defense, increased production of "pathologic" mucus and reduced the ability of hair-like cilia on the cells to remove pollutants and microorganisms.
"By sequencing RNA of almost 2,000 individual cells in the airway, we were able to present a more nuanced view of the pathology in the asthmatic airway," said Max A. Seibold, Ph.D., associate professor of Pediatrics and the Center for Genes, Environment & Health. "We found a dramatic reduction in airway defense, loss of ciliary function and a novel secretory cell that produces a thick, adhesive mucus. This provides a mechanistic explanation for the increased vulnerability of allergic asthmatics to respiratory infections that exacerbate disease."
The signaling protein IL-13 is a hallmark driver of allergic asthma, also known as type 2-high or T2-high asthma, the most common form of the disease. Dr. Seibold and his colleagues added IL-13 to cultures of airway epithelial cells derived from the trachea of two human donors. They then applied a novel technology, single-cell RNA-sequencing, to 1,025 IL-13-stimulated cells and 869 cells that did not receive IL-13. Analysis of those RNA sequences allowed the researchers to identify all the distinct cell types in the cultures and all active genes in each of those cell types.
The researchers found that the number of ciliated cells in the airways were reduced by 26 percent. Cilia are small hair-like structures whose sweeping motion moves mucus laden with particles and microorganisms up the airway and out of the body.
In addition, they found cells that normally secrete healthy mucus, various antioxidants and other defensive proteins were transformed into a novel type that produces a thick, adhesive mucus containing few defensive proteins.
"The mucus was so thick that the remaining cilia beat more slowly and were ineffective at sweeping particles away," said Nathan Jackson, Ph.D., lead author of the paper published in Cell Reports. "These results show that IL-13 transforms the airway, reducing its ability to defend against infections that cause the vast majority of asthma exacerbations."
The researchers identified 980 unique proteins in the IL-13 stimulated cultures, which confirmed that the RNA was indeed producing a dramatically altered mix of proteins. Also, the RNA sequenced in the cultured cells were strikingly similar to RNA obtained from the nasal epithelium of 441 asthmatic children, providing additional evidence that RNA sequences in the cell cultures accurately reflected processes occurring in airways of humans with allergic asthma.
The findings suggest potential therapeutic strategies to "reverse-remodel" the airway epithelium by targeting pathways controlling the IL-13 induced mucus metaplasia and other pathologic cellular differentiation, as well as reversing the transformative events underlying mucus obstruction and loss immunity.
More information: Nathan D. Jackson et al. Single-Cell and Population Transcriptomics Reveal Pan-epithelial Remodeling in Type 2-High Asthma, Cell Reports (2020). DOI: 10.1016/j.celrep.2020.107872