Computer model shows how COVID-19 could lead to runaway inflammation
A study from the University of Pittsburgh School of Medicine and Cedars-Sinai addresses a mystery first raised in March: Why do some people with COVID-19 develop severe inflammation? The research shows how the molecular structure and sequence of the SARS-CoV-2 spike protein—part of the virus that causes COVID-19—could be behind the inflammatory syndrome cropping up in infected patients.
The study, published this week in the Proceedings of the National Academy of Sciences, uses computational modeling to zero in on a part of the SARS-CoV-2 spike protein that may act as a "superantigen," kicking the immune system into overdrive as in toxic shock syndrome—a rare, life-threatening complication of bacterial infections.
Symptoms of a newly identified condition in pediatric COVID-19 patients, known as Multisystem Inflammatory Syndrome in Children (MIS-C), include persistent fever and severe inflammation that can affect a host of bodily systems. While rare, the syndrome can be serious or even fatal.
The first reports of this condition coming out of Europe caught the attention of study co-senior author Moshe Arditi, M.D., director of the Pediatric Infectious Diseases and Immunology Division at Cedars-Sinai and an expert on another pediatric inflammatory disease—Kawasaki disease.
Arditi contacted his long-time collaborator, Ivet Bahar, Ph.D., distinguished professor and John K. Vries Chair of computational and systems biology at Pitt School of Medicine, and the two started searching for features of the SARS-CoV-2 virus that might be responsible for MIS-C.
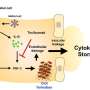
Bahar and her team created a computer model of the interaction between the SARS-CoV-2 viral spike protein and the receptors on human T cells, the foot soldiers of the immune system. Under normal circumstances, T cells help the body fight off infection, but when these cells are activated in abnormally large quantities, as is the case with superantigens, they produce massive amounts of inflammatory cytokines—small proteins involved in immune system signaling—in what's known as a "cytokine storm."
Using this computer model, the team was able to see that a specific region on the spike protein with superantigenic features interacts with T cells. Then, they compared this region to a bacterial protein that causes toxic shock syndrome and found striking similarities in both sequence and structure. Importantly, the proposed SARS-CoV-2 superantigen showed a high affinity for binding T cell receptors—the first step toward touching off a runaway immune response.
"Everything came one after another, each time a huge surprise. The pieces of the puzzle ended up fitting extremely well," said Bahar, co-senior author on the study.
By finding protein-level similarities between SARS-CoV-2 and the bacterial structure that causes toxic shock syndrome, the researchers said they may have opened up new avenues for treating not only MIS-C patients, but also adults with COVID-19 infection experiencing cytokine storm.
The researchers also collaborated with scientists studying adult COVID-19 patients in Germany and found that those who experienced severe symptoms had a T cell response similar to what is seen in people exposed to superantigens and very different from the T cell response in patients who had only mild symptoms.
"Our research finally begins to unravel the potential mechanisms involved and raises the possibility that therapeutic options for toxic shock syndrome, such as intravenous immunoglobulin and steroids, may be effective for managing and treating MIS-C in children and hyperinflammation in adult coronavirus patients," said Arditi, professor of pediatrics and biomedical sciences at Cedars-Sinai.
Arditi's and Bahar's labs are now using the ideas generated by this study to search for and test antibodies specific to the SARS-CoV-2 superantigen, with the goal of developing therapies that specifically address MIS-C and cytokine storm in COVID-19 patients.
More information: Mary Hongying Cheng et al. Superantigenic character of an insert unique to SARS-CoV-2 spike supported by skewed TCR repertoire in patients with hyperinflammation, Proceedings of the National Academy of Sciences (2020). DOI: 10.1073/pnas.2010722117