Switching off the 'survival protein' for cancer cells
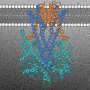
![Fig. 1: Design of peptide-modified supramolecular tweezers. a Representation of Survivin’s dimer interface based on PDB-ID: 1XOX [https://www.rcsb.org/structure/1xox]38. Both monomers, depicted in blue and gray, mainly interact via the ELTL sequence (contact region of both monomers overlapping with the NES, represented in cyan and green). This sequence was chosen as second binding motif for the peptide-modified tweezer molecules. b Representation of TW-ELTL (shown in d) bound to Survivin. TW-ELTL (yellow) binds the anchor lysine residue K103 (violet) on Survivin’s surface while the peptide motif ELTL (yellow) interacts with the ELTL region of the Survivin monomer (cyan). This is the same region of the dimer interface represented in Fig. 1a, overlapping with the NES (cyan). The chemical structures of the unmodified tweezer molecule TW (c), an asymmetrical tweezer molecule linked to the short peptide ELTL (TW-ELTL) (d), and an asymmetrical tweezer molecule linked to the elongated peptide ELTLGEFL (TW-ELTLGEFL) (e) are depicted. Credit: Nature Communications (2021). DOI: 10.1038/s41467-021-21753-9 Switching off the 'survival protein' for cancer cells](https://scx1.b-cdn.net/csz/news/800a/2021/switchingoff.jpg)
Proteins control almost all vital processes in our cells. If they do not function correctly, if there is too much or too little of them, this can lead to the development of a variety of diseases including cancer. The associated proteins are therefore also important targets for drug discovery in biomedical research.
However, there are large number of proteins that simply do not offer suitable targets for a conventional active ingredient to dock onto. That's why scientists in UDE's Collaborative Research Center 1093 are developing small, unusual molecules called supramolecular ligands that can precisely bind to their surface.
Molecules block pivotal site of the protein
Recently, the team of scientists led by Prof. Shirley Knauer, Prof. Elsa-Sanchez-Garcia and Prof. Thomas Schrader succeeded in targeting a critical interface that is important for the survival of cancer cells with such customized molecules. "The protein survivin is actually hardly found in healthy adult organisms," so Prof. Shirley Knauer. "In cancer cells, however, its production is ramped up." Using an artificial, tailored ligand, the scientists were able to cover the exact site of the survivin, which is responsible for its activation and transport out of the cell nucleus.
Each protein has a unique three-dimensional structure with a fissured surface that can form loops and niches. Sanchez-Garcia and her team performed computational analyses of the protein's surface and found out that the important interface is on an ordered but somewhat dynamic loop. Using this information and following further structural analyses, the chemists led by Schrader were able to design the ligand for this particularly difficult surface.
More information: Annika Meiners et al. Specific inhibition of the Survivin–CRM1 interaction by peptide-modified molecular tweezers, Nature Communications (2021). DOI: 10.1038/s41467-021-21753-9