Immune genetics and previous common cold infections might help protect Japan from COVID-19
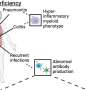
Protective immune memory—through B cells, which make antibodies, and/or T cells, which in the case of CD8+ T cells can kill virus-infected cells—can be induced by identical but also by related viruses. Related to the COVID-19 virus SARS-CoV-2, there are four common cold coronaviruses (CCCoVs) that together cause ~20% of common cold infections: OC43, HKU1, 229E, and NL63. Most adults have been infected with CCCoVs multiple times in their lives. Whether or not meaningful CCCoV-induced anti-SARS-CoV-2 neutralizing antibodies exist remains a matter of debate. Meanwhile, the generation of T cell memory should depend on the genetic make-up of the infected individual. Namely, immune recognition by T cells depends on the presentation of peptides ("epitopes") by polymorphic major histocompatibility complex (MHC) molecules, and different MHC alleles (variants between individuals) present different peptides from the same pathogen.
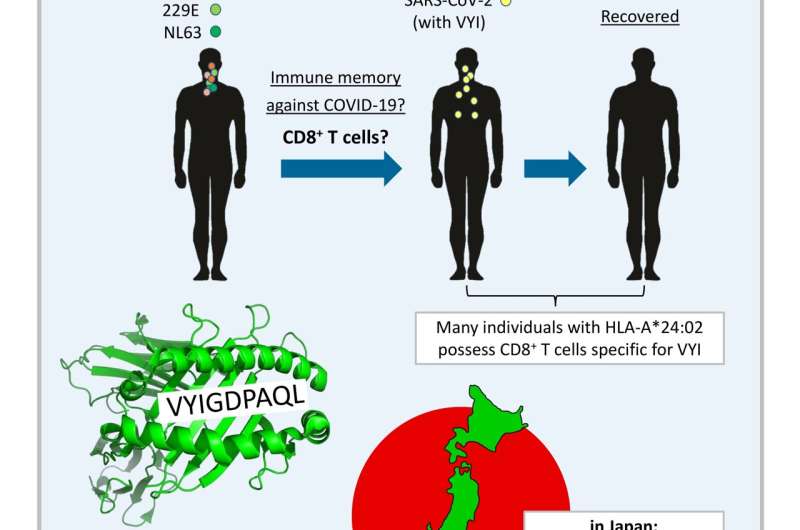
In their recent publication in F1000Research, Dijkstra, Frenette, and Dixon provided the first systematic search for SARS-CoV-2 T cell epitope sequences that were experimentally determined in one or more of >15 studies and are identical to at least one of the four CCCoVs. They found that such epitopes are rare. The only such epitope that was repeatedly identified as immunogenic by independent research groups was the peptide VYIGDPAQL, which is also named "VYI" after its first three amino acids. This peptide is a fragment of the virus helicase and identical between SARS-CoV-2 and the CCCoVs OC43 and HKU1. The VYI peptide was found to bind MHC class I allele HLA-A*24:02 and to efficiently stimulate CD8+ T cells from at least some—and sometimes the majority—of investigated HLA-A*24:02+ COVID-19 (convalescent) patients in each study that addressed this matter. This finding was confirmed in two recent studies that were not part of the Dijkstra et al. analysis.
For explaining these findings, Dijkstra et al. conservatively assumed that infections with OC43 or HKU1 had primed the anti-VYI T cells, which then were restimulated upon infection with SARS-CoV-2. The authors also speculated that the surprisingly low prevalence of COVID-19 in Japan may in part be explained by such anti-VYI T cell immune memory, since in this country ~60% of the individuals carry the allele HLA-A*24:02 and in recent years there were OC43 outbreaks. However, it is important to realize that, for now, this remains speculation only, since (i) experimental evidence for CCCoV-induced anti-VYI memory still needs to be provided, and (ii) immune memory does not necessarily mean immune protection. Nevertheless, currently, the VYI/HLA-A*24:02/Japan hypothesis put forward by the authors is one of the most plausible models for genetically explaining differences in resistance against COVID-19 between populations. Dijkstra et al. described how even a small impact on virus replication levels may have a significant impact on the spread of the virus through a population. Their article is also an interesting read on the COVID-19 situation in Japan.
Furthermore, Dijkstra et al. pointed out that the similarity between the spike (S) proteins of SARS-CoV-2 and CCCoVs is low, which suggests that the sequence conservation of this protein is relaxed. Accordingly, the authors subscribe to the fear as expressed by others that SARS-CoV-2 may mutate its S protein to escape the immune protection that is induced by the current generation of S-only vaccines. The inclusion in future vaccines of parts of the virus that are more highly conserved between SARS-CoV-2 and CCCoVs may reduce the risk of such escape. For populations in which HLA-A*24:02 is prevalent, like in Japan, the VYI peptide is a prime candidate for such inclusion.
More information: Johannes M. Dijkstra et al, Most Japanese individuals are genetically predisposed to recognize an immunogenic protein fragment shared between COVID-19 and common cold coronaviruses, F1000Research (2021). DOI: 10.12688/f1000research.51479.1