Targeting radiation resistance: Why some tumors are so stubborn

Radiation therapy has been—and will be—a cornerstone of cancer treatment for good reason: It works.
Mostly.
Currently, more than half of cancer patients receive radiation as part of their treatment. But 20 percent of them, give or take, will find that they need different options because their tumors are resistant to radiation therapy. It's a bad place to be: They may still face the potential side effects without the therapeutic benefit, and they've lost precious time.
What if clinicians had a way to predict and possibly improve radiosensitivity for individual patients? A team of researchers at the Georgia Institute of Technology and Emory University is working on something with that ultimate goal in mind.
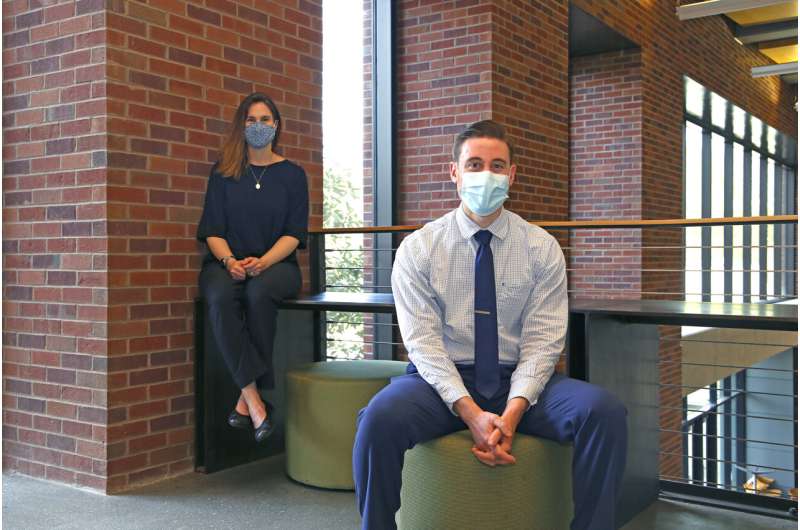
"There still isn't a great understanding of why some tumors don't respond well to radiation, and it's a significant hurdle to the long-term survival of many patients," noted Joshua Lewis, who sought answers to the radiation resistance question while a graduate student in the lab of Melissa Kemp in the Wallace H. Coulter Department of Biomedical Engineering.
Together, they've taken steps to begin to understand the underlying metabolism and build a tool to predict whether specific tumors will be one of the stubborn ones that doesn't respond. In back-to-back papers, with Lewis as lead author, Kemp said they created a new pipeline, "in which you can automatically take data, plug it into our whole cell modeling of metabolism, and actually predict the way certain tumors of various cancer types, from various patients, are going to respond."
She added: "This is the first example of really asking, with respect to radiation resistance, why there are differences that manifest themselves in tumor metabolism."
In January, they published the first study with collaborators from Wake Forest and Indiana University in the journal Cell Systems. The newest research appears this month in Nature Communications. Lewis based the studies on his Ph.D. thesis, "Genome-Scale Modeling of Redox Metabolism and Therapeutic Response in Radiation-Resistant Tumors," which he defended a year ago.
Shortly after Lewis began grad school, Wake Forest researcher Cristina Furdui approached the Kemp lab with the radiation resistance problem, "asking us to apply our expertise in systems biology," he said. "It's a more holistic approach than working with individual molecules or proteins, taking into account many different factors and their interactions, seeing if that leads to a particular response to radiation therapy."
Lewis used a well-established type of cellular modeling called "flux balance analysis," in which, "you try to model the entire metabolism of a cell—all the different chemical reactions. We model them using different biochemical equations." The researchers then plug those equations into a computer. Within seconds, they can accurately analyze about 13,000 different metabolic reactions.
"We came up with our own approach for making more accurate flux balance analysis models by integrating multiple different types of omics data," said Lewis, now pursuing his medical degree in the Emory MD/Ph.D.Program. "Omics" measure characteristics of molecules like genes, proteins, or metabolites, which comprise the cells of an organism.
By integrating genomics, transcriptomics, and metabolomics data, the researchers could model redox metabolism—the process of oxidation and reduction reactions, or the loss and gain of electrons—in cancer cells, "and use that to accurately predict how certain tumors react to radiation therapy," said Lewis, who mined data from The Cancer Genome Atlas (TCGA).
"We looked for metabolic enzymes or metabolic targets where, if you tweaked them, it could affect a tumor's radiosensitivity," Lewis said. "So, imagine if you're giving a patient radiation therapy, and you could also give them a chemotherapeutic at the same time that inhibits the action of a particular enzyme to make a tumor more sensitive to radiation."
That is, essentially, the first paper. In the second—written only by Lewis and Kemp—the researchers integrated machine learning with genome-scale metabolic modeling, "to see if we can better predict what sort of biological features are associated with a patient's response to radiation."
One main challenge for the researchers in either paper, Lewis said, was in the datasets they used from TCGA. They had good genomics and transcriptomics data, but their metabolomics data was incomplete. The computational models they developed for the Cell Systems paper helped fill in the blanks, making metabolomic predictions. They fed the data from those models into a machine learning model to better identify biomarkers of radiation resistance.
"We can think clinically of a patient giving a blood sample, and from that blood you're able to measure the levels of different metabolites and determine if the patient would be a good candidate for radiation therapy, or whether we should go ahead and think of other therapies," Lewis said.
That speaks directly to what Lewis is training for now in his M.D. program. He wants to be a pathologist and better understand how patients respond to different therapies.
"I'd like to help bridge the gap between research and the clinic," he said.
He's off to a good start, and it's exciting for Kemp.
"Josh's computational platform turns easy-to-acquire data into a model representation of hard-to-acquire attributes, like metabolic fluxes and metabolite changes, that are otherwise very challenging to measure with the scale it takes to cover many different patients," she said.
More information: Joshua E. Lewis et al. Personalized Genome-Scale Metabolic Models Identify Targets of Redox Metabolism in Radiation-Resistant Tumors, Cell Systems (2021). DOI: 10.1016/j.cels.2020.12.001
Lewis, J.E., Kemp, M.L. Integration of machine learning and genome-scale metabolic modeling identifies multi-omics biomarkers for radiation resistance. Nat Commun 12, 2700 (2021). doi.org/10.1038/s41467-021-22989-1