Tool helps predicts who will respond best to targeted prostate cancer therapy
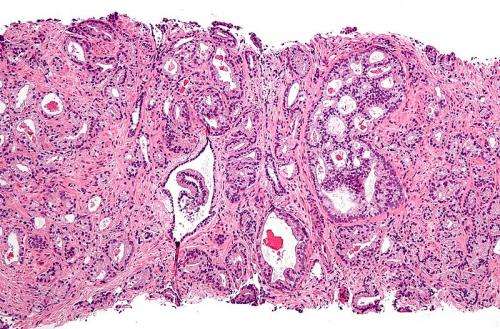
A new prognostic tool developed by researchers from the UCLA Jonsson Comprehensive Cancer Center and five other institutions helps predict which men with advanced metastatic prostate cancer will respond favorably to a novel targeted therapy.
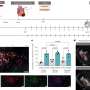
The tool, described in a study published today in Lancet Oncology, analyzes a wide spectrum of imaging and clinical data and is intended to assist physicians considering treating patients with Lutetium-177 prostate-specific membrane antigen, or LuPSMA.
LuPSMA, which binds to PSMA proteins and delivers targeted radiation to prostate cancer tissue, offers a new option to men with PSMA-positive metastatic cancer that is castration-resistant, meaning it has stopped responding to hormone therapy. LuPSMA is currently pending approval by the U.S. Food and Drug Administration.
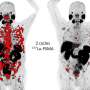
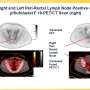
Candidates for the therapy are typically screened using a technique pioneered by UCLA and UC San Francisco called PSMA PET imaging, which combines positron emission tomography with a PET-sensitive drug to detect prostate cancer throughout the body and verify PSMA-expression in the tumors.
The new research demonstrates that a combination of clinical characteristics and PSMA PET imaging characteristics and can be used to predict which patients will have improved progression-free survival (slower disease progression) and improved overall survival (life expectancy) as a result of the treatment.
"Until now, there has been no validated tool to adequately predict the response of patients with advanced metastatic prostate cancer to LuPSMA treatment," said lead author Dr. Andrei Gafita, a postdoctoral scholar in the Ahmanson Translational Theranostics Division of the department of molecular and medical pharmacology at the David Geffen School of Medicine at UCLA. "Most important, this study suggests that screening with PSMA PET imaging could help to select patients who are most likely to benefit from this treatment."
Creating the 'nomogram'
The predictive tool—commonly called a nomogram—was developed by researchers from institutions across Europe, Australia and the U.S. who analyzed data from 270 prostate cancer patients who underwent LuPSMA treatment in clinical trials or via compassionate use.
The research team used eight parameters—including PSMA PET parameters and factors such as the time since initial diagnosis, chemotherapy status and hemoglobin levels—to create the mathematical formula that allows the tool to predict survival.
Based on the nomogram, the researchers also created a web-based risk calculator that forecasts the probability of overall and progression-free survival in response to LuPSMA. The predictions are used to stratify men into either high-risk or low-risk groups.
In the men studied, those identified by the tool as low risk had longer overall survival (24 months) and progression-free survival (6 months) than those classified as high risk (6 months and 2 months, respectively).
The findings from the research are encouraging, Gafita said, and may provide the foundation for patient selection for LuPSMA therapy. Nevertheless, he notes, until its clinical validity is demonstrated in prospective trials, the tool should be used cautiously and should not replace the clinical judgement of treating physicians.
"Our validated tool can find application particularly at institutions where LuPSMA is just being introduced as a novel therapeutic option," he said.
More information: Lancet Oncology (2021). DOI: 10.1016/S1470-2045(21)00274-6