Approach predicts novel 'protein partners' that could contribute to COVID-19 symptoms
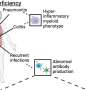
COVID-19 not only causes symptoms characteristic of a typical respiratory disorder, but has also been known to trigger a wide range of other symptoms in people who had been infected, some lasting even long after individuals test negative for the virus. These symptoms can include abnormal blood clotting, heart damage and failure, kidney disease, brain fog (confusion, memory loss, or difficulty focusing), gastrointestinal problems, and even male infertility.
Yet the mechanisms by which COVID-19 causes these diverse complications remain poorly understood.
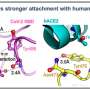
In a new paper published in the journal PeerJ, John (Jack) Werren, the Nathaniel and Helen Wisch Professor of Biology at the University of Rochester, and recent undergraduates Austin Varela '20 and Sammy Cheng '21 studied proteins that closely evolve with Angiotensin-converting enzyme 2 (ACE2), the receptor used by the SARS-CoV2 virus to enter human cells.
Using an evolutionary approach, the researchers detected proteins that "coevolve" with ACE2 in mammals as a way to identify networks of proteins that likely interact with ACE2 during its normal functions in the human body. Their rationale is that disruptions caused by COVID-19 in these normal functions of ACE2 could contribute to the unusual pathologies of the disease.
COVID-19, ACE2, and protein partners
The method used by the researchers revealed a number of candidate protein partners for ACE2 that have not previously been identified as ACE2 interactors, but which could have direct bearing on the complications experienced by people infected with the virus. These COVID-related complications can include excessive blood clotting as well as an overactive inflammation response known as a "cytokine storm"—both of which can cause tissue and organ damage.
For example, one hallmark of severe COVID-19 is abnormal blood coagulation throughout the body. The team's research revealed noteworthy connections between ACE2 and key proteins involved in the coagulation pathway. Another protein, Clusterin, which plays a significant role in "quality control" in the blood by removing misfolded proteins, strongly coevolves with ACE2—implying that they interact with each other biologically. Several proteins involved in cytokine signaling appear to coevolve with ACE2 as well.
"We propose that ACE2 has novel protein interactions that are disrupted during SARS-CoV-2 infection, contributing to the spectrum of COVID-19 pathologies," Werren says. Finding that ACE2's evolutionary partners are involved in blood coagulation and cytokine signaling is consistent with this possibility.
"These candidate protein interactions will need to be validated," Werren says. "But if supported, our findings could inform development of better treatments and therapeutics for COVID-19 and chronic complications that may arise."
As an evolutionary geneticist, Werren's research focuses on the interaction of genomes in symbiotic or parasitic relationships. When the COVID-19 pandemic hit, Werren received a grant from the National Science Foundation's Rapid Response Research program to study the ACE2's protein interactions and its network of coevolving proteins. The University's Nathaniel and Helen Wisch Chair Research Fund, meanwhile, helped support Varela's and Cheng's participation.
"Working on this project was a great opportunity for me," says Varela, the study's first author, who started researching protein-protein interactions in the Werren Lab during his sophomore year at Rochester. "Once we uncovered the evolutionary rate partners of ACE2, their potential clinical relevance was immediately clear."
Evolutionary approach to detecting protein partners and interactions
Werren and his colleagues used a computational and evolutionary approach called evolutionary rate correlation (ERC). The underlying concept is that proteins with similar rates of change during evolution are more likely to have functional interactions. For example, if you look at a phylogenetic tree depicting the evolutionary relationships among mammals, when one protein evolves quickly in a particular species, a protein with which it functionally interacts will tend to evolve quickly as well, and vice versa.
Werren previously applied the ERC method to protein interactions involved in mitochondrial function. Mitochondria are cellular structures that—among other functions—produce energy for the cell. In that study, the ERC method accurately predicted nuclear encoded proteins known to interact with mitochondria, and also detected proteins not previously known to have mitochondrial function.
Biomedical researchers currently harness a number of powerful tools for detecting and exploring proteins that interact with each other in biological processes. These include genetic screens, protein coprecipitation, and proteomic profiling.
Evolutionary approaches such as ERC have the potential to become useful additions to the biomedical research toolkit. Werren and his team are now expanding the number of proteins in their analysis. He says, "We hope to produce a more extensive database for researchers so they can explore coevolving partners for proteins of interest in their research."
More information: Austin A. Varela et al, Novel ACE2 protein interactions relevant to COVID-19 predicted by evolutionary rate correlations, PeerJ (2021). DOI: 10.7717/peerj.12159