Researchers discover that proteasome molecules can join forces to prevent cancer
Managing the waste that cells produce is an essential function of the human body, as any defect in its elimination mechanisms can lead to cancer and neurodegenerative diseases. Now a Canadian study sheds light on a new mechanism of action of the cellular systems responsible for waste elimination.
Published in Nature Communications, the study was conducted by a research team led by Université de Montréal medical professor El Bachir Affar at the UdeM-affiliated Maisonneuve-Rosemont Hospital Research Center.
Affar's team looked at a tiny biological machine called the proteasome that exists in every cell in the body. This machine is responsible for breaking down and removing unwanted, malformed or excess proteins, a vital process for the proliferation and normal functioning of the cells.
In addition, this process ensures the recycling of amino acids, which are the basic building blocks that the cells use to make new proteins. However, the exact mechanisms underlying the normal function of the proteasome are still poorly understood.
A mechanism of cell suicide
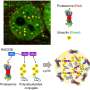
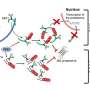
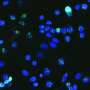
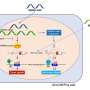
In their lab, Affar's team discovered that under conditions of cellular stress such as nutrient deprivation, proteasome molecules assemble together and form large structures called bodies or foci, which acquire new functions by inducing cell death through a mechanism of cell suicide called apoptosis.
This mechanism appears to be important in maintaining the proper functioning of tissues and organs.
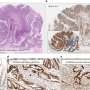
In addition, the researchers showed that this process becomes abnormal in tumoral cells, which establishes its importance in preventing the development of cancer. The new study, therefore, opens up many perspectives in understanding the functioning of normal cells and cancer cells.
"This discovery is very exciting," said Maisonneuve-Rosemont oncologist Dr. Pierre Dubé. "Dr. Affar and his team have discovered a new process that when faulty, can contribute to cancer. This opens the way to a new field of study and could lead to the identification of molecular targets for the treatment of cancers."
Added Affar, "Joining forces applies to proteasomes that unite to accomplish their mission on a microscopic scale, of course, but it turns out that this logic is also spontaneously applied at the scale of research teams and the scientific process."
'A beautiful multidisciplinary collaboration'
He cited the "beautiful multidisciplinary collaboration" between his UdeM colleagues Przemyslaw Sapieha, Mikhail Sergeev, Benjamin H. Kwok, Laura Hulea, Frédérick A. Mallette, Éric Milot, Bruno Larrivée and Hugo Wurtele, along with Jean-Yves Masson of Université Laval.
With a grant he recently received from the Canadian Institutes of Health Research, Affar plans to determine why and how proteasome orchestrates its assembly into foci. His team will use cutting-edge tools and approaches to understand the biological function and mechanism of action of proteasome foci.
His team's research will help improve scientists' knowledge of the mechanisms of proteasome function in normal cells, he hopes. By highlighting the defects that cause the proteasome to function abnormally in disease, better therapeutic targets for clinical interventions to treat cancer could be found, he said.
More information: Maxime Uriarte et al, Starvation-induced proteasome assemblies in the nucleus link amino acid supply to apoptosis, Nature Communications (2021). DOI: 10.1038/s41467-021-27306-4