Specific metabolic dependencies of cancer cells revealed by perturbation with tailored chemical library
Many types of cancer exhibit changes in their cellular metabolism. These contribute to the development and progression of cancer. Metabolic reprogramming has thus been recognized as a hallmark of cancer and may represent a vulnerability to be exploited by targeted cancer therapy. Scientists from the research group of Giulio Superti-Furga, scientific director at the CeMM Research Center for Molecular Medicine of the Austrian Academy of Sciences and Professor at the Medical University of Vienna, have now used a drug library of 243 compounds targeting a variety of metabolic pathways to identify sensitivities among 15 myeloid leukemia cell lines. They were able to identify several specific pharmacological interventions possibilities.
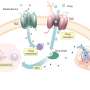
In order to grow and duplicate rapidly, proliferating cancer cells adapt their metabolism to meet the increased bioenergetic and biosynthetic demand. In fact, altered metabolism is considered a cancer hallmark. Cancer cells depend on this high-powered metabolic state to survive and grow, both in the body but also in cell culture. For some years now, the laboratory of Giulio Superti-Furga's at CeMM and MedUni Vienna has been working on understanding the dependency of specific functions in human cells on metabolites and nutrients. In a new study published in the journal Nature Communications, they report on the use of a small chemical compound library, called CLIMET, for CeMM Library of Metabolic Drugs, for the purpose of experimentally testing which part of the altered metabolic program is most important, and thus critical, to different cancer cell types. The library consists of 243 active ingredients that influence the metabolism of cells by acting on different branches of the large, intricate and widely connected network underlying cellular metabolism. The results highlight specific metabolic vulnerabilities of certain leukemia cell types, that may help conceive new therapeutic approaches.
Drug sensitivity provides important clues for therapeutic approaches
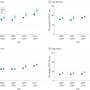
During her postdoc in the Superti-Furga laboratory at CeMM, first author Tea Pemovska developed the metabolic drug library, carefully selected for substances targeting individual pathways across the broad spectrum of metabolic processes operating in human cells. To get a better understanding of the molecular processes involved in cancer cell metabolism, the scientists performed a proof-of-concept survey using CLIMET on various blood cancer cell lines as well as patient samples. The obtained drug sensitivity profiles allowed the stratification of myeloid leukemia cell lines in five functional groups, each defined by differential sensitivity to 18 different compounds.
Study leader Giulio Superti-Furga says, "The collection of chemical agents that affect different aspects of cancer metabolism provides a toolkit to functionally assess cell lines, primary samples from cancer patients, and animal models in a versatile and dose-dependent way for their particular dependence on metabolic processes. Through this, we can get stratify cancer cell types not only by their molecular profile but by their actual metabolic needs. It is just a showcase but suggests a practical and actionable path towards an approach that exploits these cancer cell dependencies and vulnerabilities therapeutically, typically in combination with other drugs."
Identifying drivers and vulnerabilities of cell metabolism
Tea Pemovska, who is currently a scientist in the Functional Precision Hematology Group at MedUni Vienna, and colleagues, were able to show that certain human leukemia cell lines were particularly sensitive to the PI3K inhibitor Pictilisib, the fatty acid synthase inhibitor GSK2194069 and the SLC16A1 inhibitor AZD3965. She says, "Some myeloid leukemia cell lines, especially two chronic myeloid leukemia cells, showed high selective sensitivity to the inhibitor AZD3965, which inhibits the important lactate transporter SLC16A1. This allows conclusions to be drawn about which cells and/or patients might best respond to this agent."
At the same time, the study highlights the usefulness of a carefully assembled drug library with a metabolic focus to be used in phenotypic screening platforms, allowing the identification of metabolic dependencies. "Our study just delineates the feasibility of the strategy and emphasizes the importance of teasing out vulnerabilities of cancer cells by functional assays" says Giulio Superti-Furga.
More information: Metabolic drug survey highlights cancer cell dependencies and vulnerabilities. Nature Communications (2021). DOI: 10.1038/s41467-021-27329-x