Rapid peptide discovery and 'plug-and-play' technology could make personalised cancer vaccines reality
Scientists have created a pipeline for identifying, prioritizing and evaluating potential tumor antigens for the fast generation of cancer vaccines, according to a report published today in eLife.
The new approach could help researchers quickly identify tumor-specific antigens recognized by cytotoxic T cells, generating a powerful, durable and highly specific response against an individual's tumor. This could lead in turn to a quicker and easier way to generate effective, personalized cancer vaccines based on the identified antigens.
"For a cancer vaccine to be effective, we need to select target antigens that elicit a strong immune response, are exclusively present on cancer cells and are tailored to an individual's unique tumor type," explains first author Sara Feola, Postdoctoral Researcher at the ImmunoViroTherapy Lab (IVTLab), University of Helsinki, Finland. "However, only a few, if any, of the antigens on a tumor meet those characteristics, making it very difficult to identify and prioritize potentially effective candidates. Our pipeline comprises all the essential steps for the optimal development of a therapeutic cancer vaccine, but which could be carried out much more quickly on an individual patient basis, enabling true personalized therapy."
"Developing personalized cancer vaccines needs several different technologies working together and working fast," adds senior author Vincenzo Cerullo, Professor of Biological Drug Development at the University of Helsinki and group leader at IVTLab. "We need fast and reliable methods to identify and prioritize antigens, as well as rapid, inexpensive and feasible approaches to deliver these antigens to patients. During the past six years, we've been working on a project supported by the European Research Council (ERC) to make all the pieces of this complex puzzle work together, creating the pipeline that has been partially described in this work.
"Our research, which builds on previous work, involves developing a novel approach to identify tumor-specific antigens from very small samples, creating a novel algorithm to prioritize peptides based on their similarity to pathogen-derived peptides, and building several different plug-and-play technologies to deliver these peptides together with viruses or bacteria that kill cancer cells."
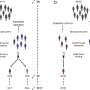
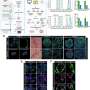
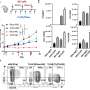
In the current study, the team began by investigating the antigen landscape of a tumor cell—that is, all the different peptides on the cell surface. They studied a mouse model of colon cancer and used state-of-the-art technologies, such as an immunopeptidomic approach based on mass spectrometry analysis, to explore surface antigens on the cell. This generated a list of thousands of peptide candidates and presented a challenge of how to prioritize them.
The team used two parallel lines of investigation: first, they looked at the relative amounts of the peptides on cancer cells compared with normal cells. This gave them clues as to whether the antigen was truly tumor specific. Second, they used a software tool previously developed in their lab to identify tumor antigens that are similar to known pathogen antigens, exploiting their potential ability to cause a similar immune response to the pathogen antigens.
Using these methods, the team narrowed their candidate list down from thousands to 26 antigen candidates. They then studied the potential of these antigens further by testing how well they stimulated T cells, and how effectively they bind to an adenoviral vector that would form the basis of the vaccine. All the candidate antigen peptides interacted with the viral vector, but six peptides performed best and were taken forward for further tests.
The next stage was to see whether a vaccine carrying these target antigens could stimulate enough of an immune response to control or halt tumor growth. To test this, the team used mice with colon tumors on their left and right flanks. They then treated one side of the mice with the vaccine coated with each of the candidate peptide antigens. As hoped, they found that vaccines carrying the peptides improved anti-tumor growth in the treated tumor, but one of the vaccines improved anti-tumor growth in the untreated tumors—suggesting that the peptide antigen in this vaccine had mounted a powerful systemic immune response against the tumors.
"We have developed and validated a pipeline that covers for the first time all the stages of personalized cancer vaccine development, starting with isolating peptides from a primary tumor to analyzing them to identify the best candidates," Cerullo concludes. "This pipeline is currently being validated in human cancer patients under our flagship project on precision cancer medicine, iCAN.
"Together, our findings demonstrate the feasibility of applying the pipeline to generate a tailored cancer vaccine by focusing on the prioritization and selection criteria and adopting quick plug-and-play technology, called PeptiCRAd, through decorating a clinically approved adenovirus vector with the selected peptides. This opens up the possibility of rapidly generating vaccines for clinical use, where effective personalized therapies represent a major goal of successful treatment."
More information: Sara Feola et al, A novel immunopeptidomic-based pipeline for the generation of personalized oncolytic cancer vaccines, eLife (2022). DOI: 10.7554/eLife.71156