Epigenetic markers predict complications in patients with type 2 diabetes
A new study by researchers at Lund University in Sweden supports the notion that patients with type 2 diabetes should be divided into subgroups and given individualized treatment. The study demonstrates that there are distinct epigenetic differences between different groups of patients with type 2 diabetes. The epigenetic markers are also associated with different risks of developing common complications in type 2 diabetes, such as stroke, heart attack and kidney disease.
"We show that there are distinct epigenetic differences between subgroups of patients with type 2 diabetes. The epigenetic markers are associated with different risks of developing common complications in diabetes, such as heart attack, stroke, and kidney disease," says Charlotte Ling, professor of diabetes and epigenetics at Lund University and lead author of the study, published in Diabetes Care.
An previous study by researchers at Lund University, published in 2018, demonstrated that it is possible to divide type 1 diabetes and type 2 diabetes into five subgroups. In November 2021, the same authors published a new study which highlighted genetic differences between the four subgroups of type 2 diabetes, suggesting different causes of the disease.
The latest study shows that there are also epigenetic differences between the four subgroups with type 2 diabetes. The epigenetic markers can be developed to predict common complications of type 2 diabetes, which would allow for tailored treatments of patients.
"Many patients with type 2 diabetes are offered standard treatments by the health care system, but growing evidence suggests that these patients need tailored treatments. Our new study adds to the evidence base that it is clinically relevant to classify patients with type 2 diabetes into subgroups to allow for more personalized treatments," says Charlotte Ling, who leads a research group in diabetes and epigenetics at Lund University.
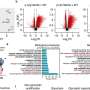
The new study encompasses 533 individuals recently diagnosed with type 2 diabetes from two population-based cohorts in Sweden. The authors measured DNA methylations in the blood at 800,000 sites in the genome of all participants. DNA methylation is a chemical process through which methyl groups attach to the DNA molecule, affecting the function of genes. The researchers found that the four subgroups had different levels of DNA methylation at 4,465 sites.
The findings were used to develop epigenetic risk scores to predict common complications of type 2 diabetes. Epigenetic markers associated with two of the subgroups could predict an increased risk of developing heart attack, stroke, and kidney disease.
"Heart attack and stroke are responsible for most deaths among patients with type 2 diabetes. Kidney disease causes a lot of suffering and is very costly for society, as many patients need dialysis treatment. An epigenetic biomarker that can predict complications at an early stage would make preventive actions possible," says Charlotte Ling.
The authors will need to verify their results in other population-based cohorts. They are also planning to study DNA methylation in tissues from, for example, muscle, adipose tissue, liver, and the pancreas of the four subgroups with type 2 diabetes.
More information: Silja Schrader et al, Novel Subgroups of Type 2 Diabetes Display Different Epigenetic Patterns, Which Associate With Future Diabetic Complications, Diabetes Care (2022). DOI: 10.2337/dc21-2489