Gene interactions classify bowel tumors for personalized medicine
Bowel cancers can be grouped into six clinically meaningful subtypes based on patterns of gene interactions commonly seen within tumor cells, according to research published today in eLife.
This new taxonomy of bowel cancer will aid understanding of the disease and could ultimately be used by doctors for precision medicine—helping them to tailor the best treatments to individual patients.
"Currently, disease staging is used to guide clinical management of people with bowel cancer, but this is inadequate because there can be substantial differences between tumors at a similar stage," explains lead author Zaoqu Lu, Master at The First Affiliated Hospital of Zhengzhou University, Zhengzhou, China.
"More detailed molecular classification of bowel cancer is more effective, but it is still based on a snapshot of the gene activity levels within a tumor at one point in time (for example, at a biopsy), and does not reflect the changing dynamics of gene activity and how genes interact with each other."
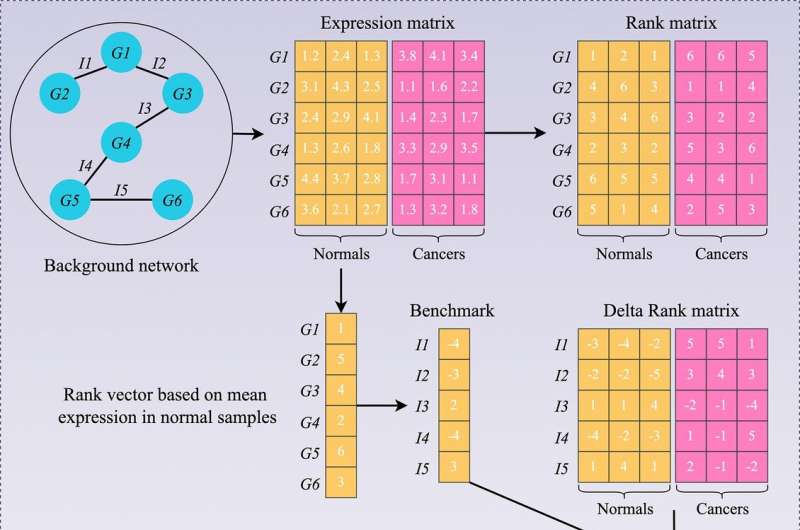
By contrast, biological interaction networks—that is, maps of which genes commonly interact with each other—remain relatively stable irrespective of time and can more reliably characterize tissues. Moreover, gene interactions are highly conserved in normal tissue but extensively altered in disease. By quantifying an "interaction change" for different gene pairs, it becomes possible to compare the overall change in gene interactions in diseased tissue, with that of the background level in normal tissue.
The team applied this gene interaction network (GIN) approach using more than 2,000 bowel cancer tissue samples and 308 normal bowel tissue samples to construct a large-scale network of changes in gene interactions. Using the network, they were then able to group bowel cancers into six subtypes, each with different clinical and molecular features. The features included important tumor characteristics, such as the proportion of cancer cells to other cell types, and whether the cancer cells were similar to stem cells, which can influence how likely a tumor is to grow and spread.
The GIN classification also identified groups with different immunogenicity—that is, how likely the bowel tumor tissue was to stimulate an immune response, which is crucial for determining whether patients might respond to immunotherapy. Other useful characteristics included likely prognosis, resistance to radiotherapy and immunotherapy treatments, and likelihood of sensitivity to different chemotherapy drugs—all of which can help tailor treatment. When the GIN approach was tested in 19 further bowel cancer datasets, the same six subtypes were consistently identified.
The first GIN was based on more than 2,000 interactions between nearly 1,400 genes, and resulted in a classifier based on 289 genes. This would be impossible to test for in a clinical setting, so to provide a rapidly accessible clinical tool, the team reduced the classifier to a randomly selected 14-gene mini-classifier. When they used the mini-classifier on a subset of 214 bowel cancer samples they found it was as stable as the full classifier at identifying the six bowel cancer subtypes.
"Our study has identified and validated a high-resolution classification system which could serve as a tool for optimizing treatment decisions for patients with bowel cancer," says senior author Xinwei Han, Professor & Director of the Department of Interventional Radiology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, China.
"The next step will be to confirm the biological and clinical interpretability of the six bowel cancer subtypes in a prospective clinical trial, but we believe that this new taxonomy could facilitate more effective personalized treatment of patients with bowel cancer."
More information: Zaoqu Liu et al, Gene interaction perturbation network deciphers a high-resolution taxonomy in colorectal cancer, eLife (2022). DOI: 10.7554/eLife.81114