This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
Validation of a comprehensive genomic profiling assay
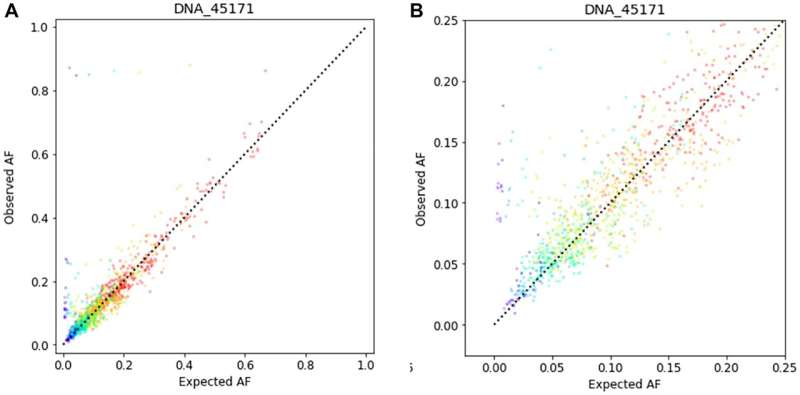
A new research paper titled "Analytic validation of NeXT Dx, a comprehensive genomic profiling assay" has been published in Oncotarget.
In this new research paper, researchers Juan-Sebastian Saldivar, Jason Harris, Erin Ayash, Manqing Hong, Prateek Tandon, Saloni Sinha, Patricia Miranda Hebron, Erin E. Houghton, Kaleigh Thorne, Laurie J. Goodman, Conan Li, Twinkal R. Marfatia, Joshua Anderson, Massimo Morra, John Lyle, Gabor Bartha, and Richard Chen from Personalis, Inc. describe the analytic validation of NeXT Dx, a comprehensive genomic profiling assay to aid therapy and clinical trial selection for patients diagnosed with solid tumor cancers.
"The NeXT Dx clinical report currently provides the ordering clinician with information from 401 cancer-related genes on clinically relevant mutations, as well as related drug response associations and a curated list of clinical trials that may be applicable to the patient," the researchers write.
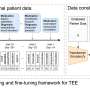
Proprietary methods were utilized to perform whole exome and whole transcriptome sequencing for detection of single nucleotide variants (SNVs), insertions/deletions (indels), copy number alterations (CNAs), and gene fusions, and determination of tumor mutation burden and microsatellite instability. Variant calling is enhanced by sequencing a patient-specific normal sample from, for example, a blood specimen. This provides highly accurate somatic variant calls as well as the incidental reporting of pathogenic and likely pathogenic germline alterations. Fusion detection via RNA sequencing provides more extensive and accurate fusion calling compared to DNA-based tests.
NeXT Dx features the proprietary Accuracy and Content Enhanced technology, developed to optimize sequencing and provide more uniform coverage across the exome. The exome was validated at a median sequencing depth of >500x. While variants from 401 cancer-associated genes are currently reported from the assay, the exome/transcriptome assay is broadly validated to enable reporting of additional variants as they become clinically relevant. NeXT Dx demonstrated analytic sensitivities as follows: SNVs (99.4%), indels (98.2%), CNAs (98.0%), and fusions (95.8%). The overall analytic specificity was >99.0%.
The researchers explain, "By more comprehensively characterizing the molecular characteristics of each patient's tumor, NeXT Dx provides personalized recommendations critical to clinical decision-making with respect to current FDA-approved drug-variant specific treatments and evolving treatment opportunities via enrollment in clinical trials."
More information: Juan-Sebastian Saldivar et al, Analytic validation of NeXT Dx™, a comprehensive genomic profiling assay, Oncotarget (2023). DOI: 10.18632/oncotarget.28490