This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
AI-based tool leverages diverse data sources for a new approach to rare disease diagnosis
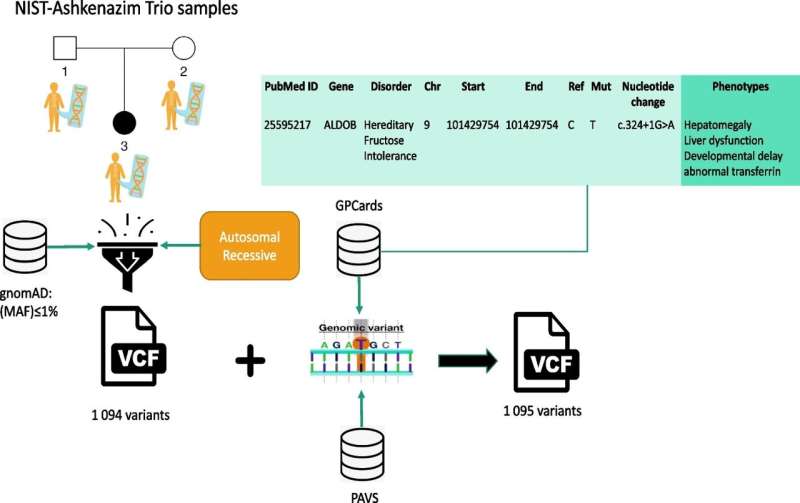
In a major step toward untangling the genetic complexities of rare diseases, KAUST researchers have unveiled an innovative AI-based tool that leverages varied symptom descriptions, along with evidence from the scientific literature and genomic datasets, to pinpoint disease-associated gene variants. This tool could aid in the diagnosis of these enigmatic conditions.
Named STARVar, the method leverages a diverse range of data sources—including background information from the scientific literature, genomic information from DNA sequence reads and clinical symptoms from individual patient records—to precisely identify genetic variants associated with diseases.
The new artificial intelligence-powered resource stands apart from other gene prioritization tools because of its focus on real-world patient symptoms, regardless of how these clinical descriptions are documented. The study is published in the journal BMC Bioinformatics.
"STARVar stands a unique and efficient tool that has the advantage of prioritizing genomic variants by using flexibly expressed patient symptoms in free-form text," says Șenay Kafkas, a bioinformatics researcher at KAUST and the first author of a new report that details the innovative tool.
Traditional methods often demand that clinical presentations adhere to standardized vocabularies, impeding a more nuanced and accurate understanding of patient symptoms. The reality, however, is that doctors and researchers frequently convey patient data using terminology that extends beyond predefined terms.
STARVar—short for Symptom-based Tool for Automatic Ranking of Variants—now offers a solution that is more dynamic and adaptable in practice.
Designed by KAUST computer scientist Robert Hoehndorf and members of his team, the method can interpret symptom data recorded in either standardized or natural language formats.
When evaluated on different genomic datasets—generated using clinical variants collected from patients, both in Saudi Arabia and from other countries around the world—STARVar outperformed several other variant prioritization tools that can operate with only rigidly represented symptoms. In particular, the algorithm consistently ranked the correct disease-associated variant at or near the top of the list of potential candidate variants in these validation tests.
Illustrating the impact of STARVar in a real-world setting, the researchers also used the tool to help diagnose a young Saudi girl who showed signs of joint stiffness, lumps under the skin and bone damage.
Out of nearly 800 suspect gene variants uncovered by genomic sequencing, STARVar deftly narrowed down the possibilities to a solitary mutation. This mutation, in a gene called MMP2, was already known to be pathogenic and thus was implicated as the likely driver of the girl's condition.
STARVar is now freely available online, and Kafkas hopes to see clinical genetics community embracing it and integrating the analytic method into their genomic workflows. "STARVar stands as a unique and efficient tool," she says, "one that will shed light on rare diseases and provide vital diagnostic support to clinicians and affected families."
More information: Șenay Kafkas et al, Starvar: symptom-based tool for automatic ranking of variants using evidence from literature and genomes, BMC Bioinformatics (2023). DOI: 10.1186/s12859-023-05406-w