This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
Largest dataset of thousands of proteins a landmark step for research into human health

The journal Nature has published the results of the world's largest and most comprehensive study on the effects of common genetic variation on proteins circulating in the blood and how these associations can contribute to disease. This unprecedented population-scale investigation of proteins, powered by turning biological samples into data from UK Biobank, will help scientists better understand how and why diseases develop, which could help drive the development of new diagnostics and treatments for a wide range of health conditions.
To develop this unique and unparalleled dataset, researchers measured the abundance of nearly 3,000 circulating proteins, many of which were previously difficult to capture, from over 54,000 participants in the UK Biobank—which has been collecting data and tracking the health of 500,000 volunteer participants enrolled between 2006 and 2010.
The study identified over 14,000 associations between common genetic variants and proteins circulating in the blood, over 80% of which were previously unknown. Scientists worldwide will be able to access the proteomic data in the coming weeks via UK Biobank.
This landmark study was carried out by the Pharma Proteomics Project, a collaboration between 13 leading biopharmaceutical companies. The team carried out analyses on the data, demonstrating the vast potential for future research using the study. These include:
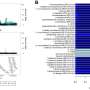
- Genome-wide association studies to build an open access library of all the common gene variants that influence protein levels in blood. This can be used to study complex biological processes, such as the immune system, find proteins that are key players in causing disease, identify new drug targets and potentially shorten development time for earlier-stage drug candidates and increase success rates for clinical trials.
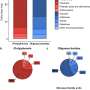
- Profiling of blood protein levels across the top 20 most common health conditions in UK Biobank. This revealed that, for example, inflammatory proteins, long thought to contribute towards mental health conditions, are significantly higher in patients with depression.
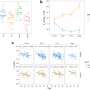
- Training machine learning models to determine how successfully blood proteins can predict demographic factors. This analysis found that blood proteins can predict age, sex and body mass index (BMI) with very high accuracy. In the future, this technology could be used to compare chronological age with biological age and determine how this is related to risk of future diseases.
Professor Naomi Allen, Chief Scientist of UK Biobank, said, "This momentous study offers whole new avenues of research to the biomedical community, and is a leading example of how cross-sector collaboration can bring about results that are so much greater than the sum of their parts."
"All of these data will soon be available to bona fide researchers across the globe, alongside the existing genomic, lifestyle and health data that UK Biobank holds for its 500,000 volunteers. I am excited for researchers to use these data to identify patterns that could transform our understanding of how diseases develop, and to identify potential new treatment pathways."
Dr. Chris Whelan, Director, Neuroscience, Data Science & Digital Health, Janssen Research & Development, LLC, a Johnson & Johnson Company, who leads the Pharma Proteomics Project, added, "To date, the scientific community has invested substantially in genomics for the advancement of precision medicine. However, to identify the right drug for the right patient at the right time, we must move beyond genomics alone."
"This dataset will help paint a much more nuanced and detailed picture of how the human genome and proteins circulating in the blood influence human health and disease—enabling biomedical researchers to identify new biological associations, find new drug targets and build blood-based diagnostics."
Other future innovative work expected to result from this study includes using proteins circulating in the blood to predict whether someone will develop a disease several years before the condition occurs, classifying diseases into distinct biological subtypes, and using proteins in the blood to predict drug efficacy and safety prior to clinical trials.
More information: Christopher Whelan, Plasma proteomic associations with genetics and health in the UK Biobank, Nature (2023). DOI: 10.1038/s41586-023-06592-6. www.nature.com/articles/s41586-023-06592-6