July 6, 2023 report
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Proteomic profile study reveals signatures for distinguishing different forms of Alzheimer's disease

Researchers at Washington University School of Medicine in St. Louis have identified proteomic changes associated with forms of Alzheimer's disease (AD). In a paper, "Proteomics of brain, CSF, and blood identifies molecular signatures for distinguishing sporadic and genetic Alzheimer's disease," published in Science Translational Medicine, the researchers identify specific and shared proteomic changes associated with sporadic Alzheimer's disease (AD) across brain tissue, cerebrospinal fluid (CSF), and blood.
The findings show the potential of proteomic analysis to distinguish between different types of AD and act as a predictive model and biomarker of disease types.
The study focused on three key categories: autosomal dominant AD linked to specific gene variants, cases involving risk variants in the TREM2 gene, and sporadic AD, which accounts for most AD cases. Sporadic AD, or "late-onset," is without a known genetic association and typically occurs after age 65.
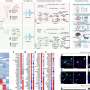
Using multiplexed, single-stranded DNA aptamer assay, the researchers analyzed 1,305 proteins across brain tissue, cerebrospinal fluid (CSF), and blood samples. They performed differential abundance analysis to identify proteins associated with AD status in each category.
The results revealed significant associations between AD status and specific proteins in the brain, CSF, and blood. In the brain, 12 proteins were significantly associated with AD status, with correlations to other AD-related traits. Six of these proteins were also detected in both CSF and blood samples and showed associations with AD status and age at onset. The findings were further validated in external datasets from various studies, confirming their reproducibility.
In CSF, the researchers identified 117 proteins significantly associated with clinical AD status. Of these, 27 proteins were found across brain tissue and blood samples. External replication studies supported the association of 40 proteins with AD status, emphasizing the potential of CSF proteomics in identifying biomarkers for AD.
Similarly, 26 proteins were associated with sporadic AD status in blood plasma, and seven proteins were replicated in brain and CSF samples. External replication studies confirmed the association of 9 proteins with AD.
The study's findings suggest that combined proteomic analysis of brain tissue, CSF, and blood plasma can provide valuable markers for distinguishing between sporadic and genetically defined AD.
Enrichment analyses highlighted several pathways related to AD, Parkinson's disease, and innate immune responses. The most strongly enriched pathways for sporadic AD were the programmed cell death and the intrinsic pathway for apoptosis, which included several proteins.
This suggests that some of the cell death associated with AD pathogenesis is regulated by apoptosis, the body's natural process of removing unrepairable cells, and not due to necrosis, entosis, ferroptosis, or lysosomal-dependent cell death.
While further validation is required, the results highlight the usefulness of integrating proteomic analysis across different tissues and fluids to comprehensively understand AD biology and develop prediction models for individuals with specific genetic profiles.
The findings offer hope for earlier diagnosis, personalized treatment strategies, and improved patient outcomes in the future.
More information: Yun Ju Sung et al, Proteomics of brain, CSF, and plasma identifies molecular signatures for distinguishing sporadic and genetic Alzheimer's disease, Science Translational Medicine (2023). DOI: 10.1126/scitranslmed.abq5923
© 2023 Science X Network