This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Findings challenge standard understanding of COVID-19 infection

Some viruses move between species. For example, SARS-CoV-2, the virus that causes COVID-19, can spill over from humans to mink, an agricultural species, and then spill back from mink to humans. Spillback is a concern because SARS-CoV-2 can mutate in the mink and come back to humans in a more virulent form. Both spillover and spillback of SARS-CoV-2 have been reported on mink farms in the United States and Europe.
To address these issues, a research team at the University of California, Riverside, has now studied zoonosis—the interspecies transmission of pathogens—in mink and found that TMPRSS2, an enzyme critical for viral fusion entry of SARS-CoV-2 in humans, is not functional in mink.
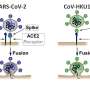
"We found mink lung cells are infected by the 'endocytosis pathway,' not the TMPRSS2 fusion pathway commonly observed in human cells," said doctoral student Ann Song, first author of the research paper, "Endocytosis inhibitors block SARS-CoV-2 pseudoparticle infection of mink lung epithelium," that appears in Frontiers in Microbiology.
"Our findings show that SARS-CoV-2 entry is not the same in all mammals and emphasize the need for thorough investigations into viral entry mechanisms across different species."
Song explained that viral fusion occurs when the membrane of the virus fuses with the plasma membrane of the host cell during infection. She said endocytosis is an essential process in which cells engulf external materials in small vesicles formed from their plasma membranes. SARS-CoV-2 can be taken up by host cells via endocytosis, she said.
"Our results show that the functional—or enzymatic—domain is missing in mink TMPRSS2," she said. "We do not know why. We think the enzyme may have multiple functions. It can do something else in mink, but it does not play a role in SARS-CoV-2 fusion to host cells. As a result, targeting TMPRSS2 would not be helpful in preventing infection in mink. What is clear is that SARS-CoV-2 entry varies among different species and tissue types."
Song said zoonosis is a public health concern as dangerous mutated forms of the virus could be introduced into the human population through spillback. During the pandemic, hundreds of papers were published on COVID-19 in humans. Now that COVID-19 in humans is under better control, scientific attention is turning to zoonosis.
Lead author Prue Talbot, a professor of the graduate division in the Department of Molecular, Cell and Systems Biology in whose lab Song works, said researchers should not underestimate the possibility of spillover and spillback of SARS-CoV-2 in other mammalian species.
"Deadly mutants can emerge from spillover/spillback events," Talbot said. "As another example, many herds of deer, which are hunted by humans, are infected with SARS-CoV-2 and are thus potential sources of spillback."
Talbot and Song were joined in the research by postdoctoral researcher Rattapol Phandthong. Next, the research team will work on the infectability of human embryos in pregnant women who have COVID-19.
To achieve their results, the researchers conducted their experiments using lung epithelial cells from mink.
More information: Ann Song et al, Endocytosis inhibitors block SARS-CoV-2 pseudoparticle infection of mink lung epithelium, Frontiers in Microbiology (2023). DOI: 10.3389/fmicb.2023.1258975