This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
CRISPR-based mapping uncovers 'switches' for immune genes central to health
Healthy T cells are the foundation of a balanced immune system. They can spring into action to fight off infections or cancer, but keep from becoming so activated that they hurt the very cells they're meant to protect, as is the case in autoimmune diseases.
Behind the scenes, a trio of genes orchestrate this careful balancing act. These genes—known as CD28, CTLA4, and ICOS—are known to play a pivotal role in shaping how T cells function and maintain their balance. They're all housed together on the same stretch of DNA, coding for proteins whose finely tuned coordination is essential for immune health.
While scientists have long recognized the vital function of these neighboring genes, much less is known about how they're switched on and off in specialized T cells that suppress the immune system. There's also a dearth of knowledge about how the switches function when T cells are in a resting state versus stimulated.
In a new study, scientists at Gladstone Institutes used a tool known as CRISPR interference, or CRISPRi, to map the layered mechanisms that control expression of these fundamental immune genes. The findings, published in Nature Genetics, provide valuable insights into immune balance, autoimmune diseases, and the development of cancer immunotherapies.
"Until recently, we lacked the tools to systematically determine the mechanistic links underlying the coordinated regulation of genes grouped as neighbors at one site in the genome," says Gladstone Senior Investigator Alex Marson, MD, Ph.D., director of the Gladstone-UCSF Institute of Genomic Immunology, who led the study.
"Now, we can begin to decipher the complex logic of a multi-gene site associated with disease risk, and we can do this directly in human cells that play a central role in those diseases."
Such knowledge is crucial for understanding the basis of immune-driven conditions. For example, a person's risk of certain autoimmune diseases may be linked to genetic variants in same DNA "neighborhood" that affect switches controlling the three immune genes. And it will help in the design of new cancer immunotherapies, which instruct a person's T cells to fight cancerous cells, Marson says.
Unlocking a new level of understanding
For Cody Mowery, an MD-Ph.D. candidate in Marson's lab and at University of California San Francisco, focusing on the stretch of DNA containing CD28, CTLA4, and ICOS was a no-brainer. The variants in this locus are known to be associated with autoimmune disorders such as lupus and inflammatory bowel disease, and some treatments targeting these genes already exist, improving and extending the lives of people with autoimmune conditions and cancer.
"I don't think it's possible to overstate how important these genes are," says Mowery, first author of the new study. "And given that they're just lying right next to one another in the genome, of course we want to go in and understand everything we can."
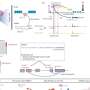
The basic controls of gene expression are well established. Some of the key players are CREs, short for "cis-regulatory elements." CREs are non-coding stretches of DNA located near the gene they regulate. CREs also interact with other regulatory elements known as trans-regulators to boost or put the brakes on DNA transcription, the first step of the gene-to-protein production line.
The Gladstone scientists wanted to discover the location of CREs and test the role that specific CREs play in regulating the three genes, which could then provide insight into underlying causes of disease risk.
"Given a certain cell type and its current state of activation, we wanted to know what gives rise to one gene being expressed versus another being suppressed in that same situation" Mowery says. "Answering that question becomes very complicated when you have highly coordinated regulation of genes in the same locus."
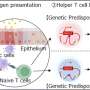
The research team used CRISPRi to disrupt more than 11,500 target sites—each in a different human T cell—across the entire stretch of DNA containing CD28, CTLA4, and ICOS. They did this in two different types of T cells from human blood donors, then examined the effects on gene expression.
"This would not have been possible before our group's recent development of methods for applying CRISPRi at scale in primary human T cells," says Jimmie Ye, Ph.D., an affiliate investigator at Gladstone and associate professor of medicine at UC San Francisco. Along with Marson, he was a co-senior author of the study.
The team's analysis—aided by the computational biology lab of Gladstone Senior Investigator Katie Pollard, Ph.D.—identified and mapped the effects of specific CREs on expression of each of the three genes under different states of cellular activation.
They found that in some cases, the expression of a particular gene is controlled by different CREs, depending on the cell type or activation state. Some CREs are shared between genes but affect each gene differently, and a CRE that affects a gene's expression might be located far from the gene itself.
"During this process, we were able to validate the functional effects of specific CREs that had previously only been indirectly associated with gene expression," Ye says.
Multi-layered complexity
Next, the researchers incorporated previously compiled datasets and additional molecular and computational tools to identify trans-regulators of the three genes and link them to the particular CREs they affect. The analysis also uncovered potential mechanisms for regulatory crosstalk between the genes and sites in the genetic region that appear to shape the 3D architecture of the DNA to make sure the switches focus their activities on the correct gene targets.
"We now have a far more dynamic picture of gene regulation at this locus," says Marson, who is also director of the Parker Institute for Cancer Immunotherapy at Gladstone Institutes. "This new level of understanding could eventually lead to better therapies for autoimmune conditions and cancer."
The team has already demonstrated one potential example: Within the identified CREs, they uncovered two genetic variants that appear to strongly affect CTLA4 expression and are associated with rheumatoid arthritis. Further research could reveal their potential clinical significance.
The researchers are also interested in the implications of their research for genome engineering to treat disease.
"There's a lot of focus on using genetic engineering to create new proteins within cells," Ye says. "But I think going forward, it's going to be about putting all the regulatory logic together in non-protein-coding sequences in ways that allow us to exert much better control."
More information: Cody T. Mowery et al, Systematic decoding of cis gene regulation defines context-dependent control of the multi-gene costimulatory receptor locus in human T cells, Nature Genetics (2024). DOI: 10.1038/s41588-024-01743-5