Cancer biology: Charting a tumor's genomic roots
Whole-genome sequencing gives researchers a deeper understanding of factors contributing to the onset and progression of gastric cancer.
By combining the skills and knowledge of a large number of cancer and genomics experts, the Singapore Gastric Cancer Consortium has successfully generated genome sequences for tumor samples from two gastric cancer patients1. The effort was coordinated by Niranjan Nagarajan, Yijun Ruan and Patrick Tan from the A*STAR Genome Institute of Singapore.
Modern DNA sequencing instruments can deliver a high-quality human genome sequence within a week for less than US$10,000. This technology offers a promising tool for revealing the specific genomic disruptions that underlie poorly understood cancers such as gastric cancer, which remains extremely challenging to diagnose and treat. Unfortunately, the repetitive sequences and large-scale chromosomal rearrangements observed in cancer genomes make it difficult to accurately assemble relatively short DNA sequence 'reads' into a complete sequence.
Nagarajan credits their success to a combination of sophisticated analytical tools. "We had deep-sequencing data from two complimentary approaches as well as a new assembly approach available to us," he says, "and these provided an ideal test-bed for carrying out this study."
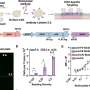
The researchers began by generating genome segments dubbed 'contigs' based on overlap between short sequencing reads. Then they used a long-range mapping technique called DNA paired-end tag analysis to accurately assemble these into even larger contigs that span a considerable portion of the genome. The resulting coverage allowed them to conduct a detailed census of small-scale sequence alterations as well as larger-scale chromosomal rearrangements. Their work revealed more than five times as many changes than could be detected with more limited conventional sequencing strategies.
Gastric cancer is unusual in that it can be triggered by infection with the bacterium Helicobacter pylori. Nagarajan and co-workers' approach allowed them to identify differences between H. pylori- and non-H. pylori-associated cancer cases. "We were able to recover the pathogen genome associated with this cancer from out of massive amounts of sequencing data," says Nagarajan. "We also found a mutational signature that can be linked to infection and is likely to have had a disproportionate impact in tumorigenesis." The researchers were further able to define mutational characteristics that might contribute to specific categories of cancer-related genomic damage, such as unnatural expansion of repetitive sequences or sizeable insertions and deletions.
This work demonstrates the valuable clinical information that tumor genome sequencing can provide. Nagarajan and co-workers are now developing better computational tools that could make better sense of the avalanche of genomic data that can be generated from a single tumor.