The genes behind the guardians of the airways
Dysfunctions in cilia, tiny hair-like structures that protrude from the surface of cells, are responsible for a number of human diseases. However the genes involved in making cilia have remained largely elusive. In the first comprehensive analysis of its kind, researchers from the A*STAR Institute of Molecular and Cell Biology in Singapore have identified hundreds of genes involved in the proper formation of a particular type of cilia that help to remove mucus and dirt from the lungs1.
"This collection of genes will be invaluable for our understanding of how cilia are made," says Sudipto Roy, a developmental biologist and senior author of the study. "More importantly, it will greatly facilitate the diagnosis of cilia-based diseases."
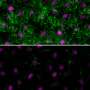
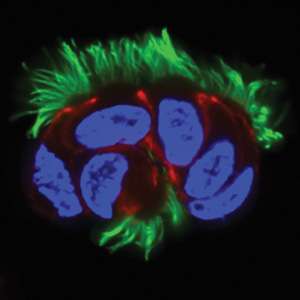
Cilia come in two forms: primary, or non-motile, cilia that serve as environmental sensors for cells throughout the body, and motile cilia, which constantly beat to sweep debris from the middle ear and respiratory tract (see image). Mutations that affect the function of motile cilia play a role in genetic disorders like primary ciliary dyskinesia, which affects the lungs and other sites where such cilia are present.
Previously, Roy and colleagues found that a transcription factor called forkhead box protein J1 (Foxj1) is the master regulator of motile cilia development2. The researchers demonstrated that Foxj1 controls the expression of numerous genes involved in the production of these moving, microscopic protrusions. However, they were unsure as to exactly which genes were implicated in the process of making functional motile cilia.
Roy's team, guided by senior research fellow Semil Choksi, therefore decided to perform a systematic analysis of all the genes in the zebrafish, a common model of cilia development. They discovered more than 500 genes with mammalian counterparts that are activated by Foxj1, the majority of which had not previously been associated with cilia production.
The researchers randomly selected 50 of these genes for functional studies. They artificially boosted expression levels of each gene by injecting lab-made RNA into zebrafish embryos, and found that around 30% of the proteins encoded by the genes localized to the motile cilia. The researchers also inactivated each of these 50 genes in turn using morpholinos, a standard knockdown tool. In this way, they showed that more than 60% of the genes were needed for the proper differentiation or functioning of motile cilia.
According to Choksi, the team's collection of cilia-associated genes is set to help researchers identify previously unknown mutations behind cilia disorders in patients—and ultimately, perhaps, new therapies.
More information: 1. Choksi, S. P., Babu, D., Lau, D., Yu, X. & Roy, S. Systematic discovery of novel ciliary genes through functional genomics in the zebrafish. Development 141, 3410–3419 (2014). dx.doi.org/10.1242/dev.108209
2. Yu, X., Ng, C. P., Habacher, H. & Roy, S. Foxj1 transcription factors are master regulators of the motile ciliogenic program. Nature Genetics 40, 1445–1453 (2008). dx.doi.org/10.1038/ng.263